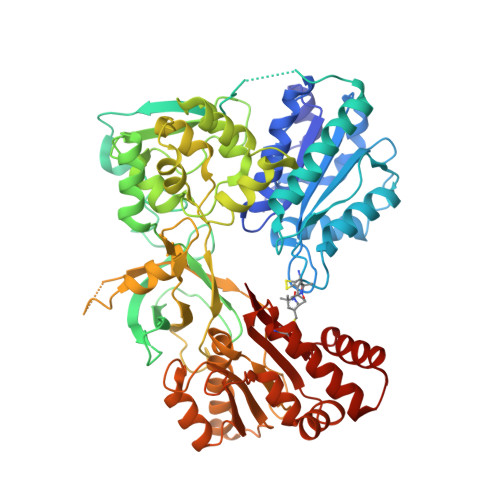
Structural and Functional Studies of the Membrane-Binding Domain of NADPH-Cytochrome P450 Oxidoreductase.
Xia, C., Shen, A.L., Duangkaew, P., Kotewong, R., Rongnoparut, P., Feix, J., Kim, J.P.(2019) Biochemistry 58: 2408-2418
- PubMed: 31009206
- DOI: https://doi.org/10.1021/acs.biochem.9b00130
- Primary Citation of Related Structures:
6NJR - PubMed Abstract:
NADPH-cytochrome P450 oxidoreductase (CYPOR), the essential flavoprotein of the microsomal cytochrome P450 monooxygenase system, is anchored in the phospholipid bilayer by its amino-terminal membrane-binding domain (MBD), which is necessary for efficient electron transfer to cytochrome P450. Although crystallographic and kinetic studies have established the structure of the soluble catalytic domain and the role of conformational motions in the control of electron transfer, the role of the MBD is largely unknown. We examined the role of the MBD in P450 catalysis through studies of amino-terminal deletion mutants and site-directed spin labeling. We show that the MBD spans the membrane and present a model for the orientation of CYPOR on the membrane capable of forming a complex with cytochrome P450. EPR power saturation measurements of CYPOR mutants in liposomes containing a lipid/Ni(II) chelate identified a region of the soluble domain interacting with the membrane. The deletion of more than 29 residues from the N-terminus of CYPOR decreases cytochrome P450 activity concomitant with alterations in electrophoretic mobility and an increased resistance to protease digestion. The altered kinetic properties of these mutants are consistent with electron transfer through random collisions rather than via formation of a stable CYPOR-P450 complex. Purified MBD binds weakly to cytochrome P450, suggesting that other interactions are also required for CYPOR-P450 complex formation. We propose that the MBD and flexible tether region of CYPOR, residues 51-63, play an important role in facilitating the movement of the soluble domain relative to the membrane and in promoting multiple orientations that permit specific interactions of CYPOR with its varied partners.
Organizational Affiliation:
Department of Biochemistry , Medical College of Wisconsin , Milwaukee , Wisconsin 53226 , United States.