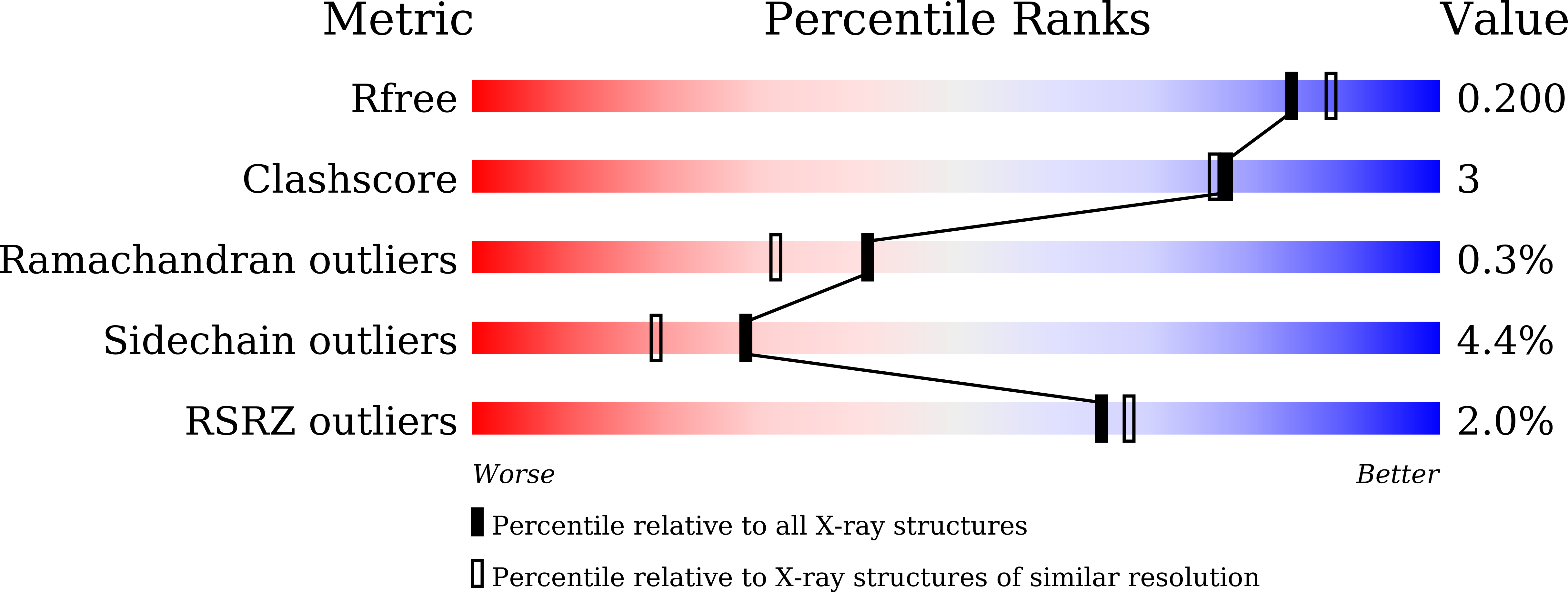
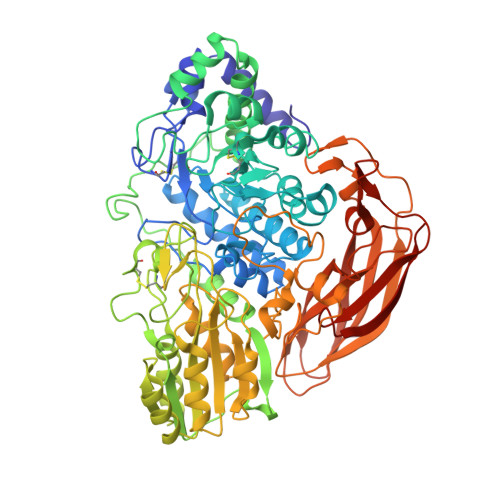
Chaetomella raphigerabeta-glucosidase D2-BGL has intriguing structural features and a high substrate affinity that renders it an efficient cellulase supplement for lignocellulosic biomass hydrolysis.
Kao, M.R., Kuo, H.W., Lee, C.C., Huang, K.Y., Huang, T.Y., Li, C.W., Chen, C.W., Wang, A.H.J., Yu, S.M., Ho, T.H.D.(2019) Biotechnol Biofuels 12: 258-258
- PubMed: 31700541
- DOI: https://doi.org/10.1186/s13068-019-1599-0
- Primary Citation of Related Structures:
6JXG - PubMed Abstract:
To produce second-generation biofuels, enzymatic catalysis is required to convert cellulose from lignocellulosic biomass into fermentable sugars. β-Glucosidases finalize the process by hydrolyzing cellobiose into glucose, so the efficiency of cellulose hydrolysis largely depends on the quantity and quality of these enzymes used during saccharification. Accordingly, to reduce biofuel production costs, new microbial strains are needed that can produce highly efficient enzymes on a large scale. We heterologously expressed the fungal β-glucosidase D2-BGL from a Taiwanese indigenous fungus Chaetomella raphigera in Pichia pastoris for constitutive production by fermentation. Recombinant D2-BGL presented significantly higher substrate affinity than the commercial β-glucosidase Novozyme 188 (N188; K m = 0.2 vs 2.14 mM for p -nitrophenyl β-d-glucopyranoside and 0.96 vs 2.38 mM for cellobiose). When combined with RUT-C30 cellulases, it hydrolyzed acid-pretreated lignocellulosic biomasses more efficiently than the commercial cellulase mixture CTec3. The extent of conversion from cellulose to glucose was 83% for sugarcane bagasse and 63% for rice straws. Compared to N188, use of D2-BGL halved the time necessary to produce maximal levels of ethanol by a semi-simultaneous saccharification and fermentation process. We upscaled production of recombinant D2-BGL to 33.6 U/mL within 15 days using a 1-ton bioreactor. Crystal structure analysis revealed that D2-BGL belongs to glycoside hydrolase (GH) family 3. Removing the N-glycosylation N68 or O-glycosylation T431 residues by site-directed mutagenesis negatively affected enzyme production in P. pastoris . The F256 substrate-binding residue in D2-BGL is located in a shorter loop surrounding the active site pocket relative to that of Aspergillus β-glucosidases, and this short loop is responsible for its high substrate affinity toward cellobiose. D2-BGL is an efficient supplement for lignocellulosic biomass saccharification, and we upscaled production of this enzyme using a 1-ton bioreactor. Enzyme production could be further improved using optimized fermentation, which could reduce biofuel production costs. Our structure analysis of D2-BGL offers new insights into GH3 β-glucosidases, which will be useful for strain improvements via a structure-based mutagenesis approach.
Organizational Affiliation:
1Molecular and Cell Biology, Taiwan International Graduate Program, Academia Sinica and National Defense Medical Center, Taipei, Taiwan, ROC.