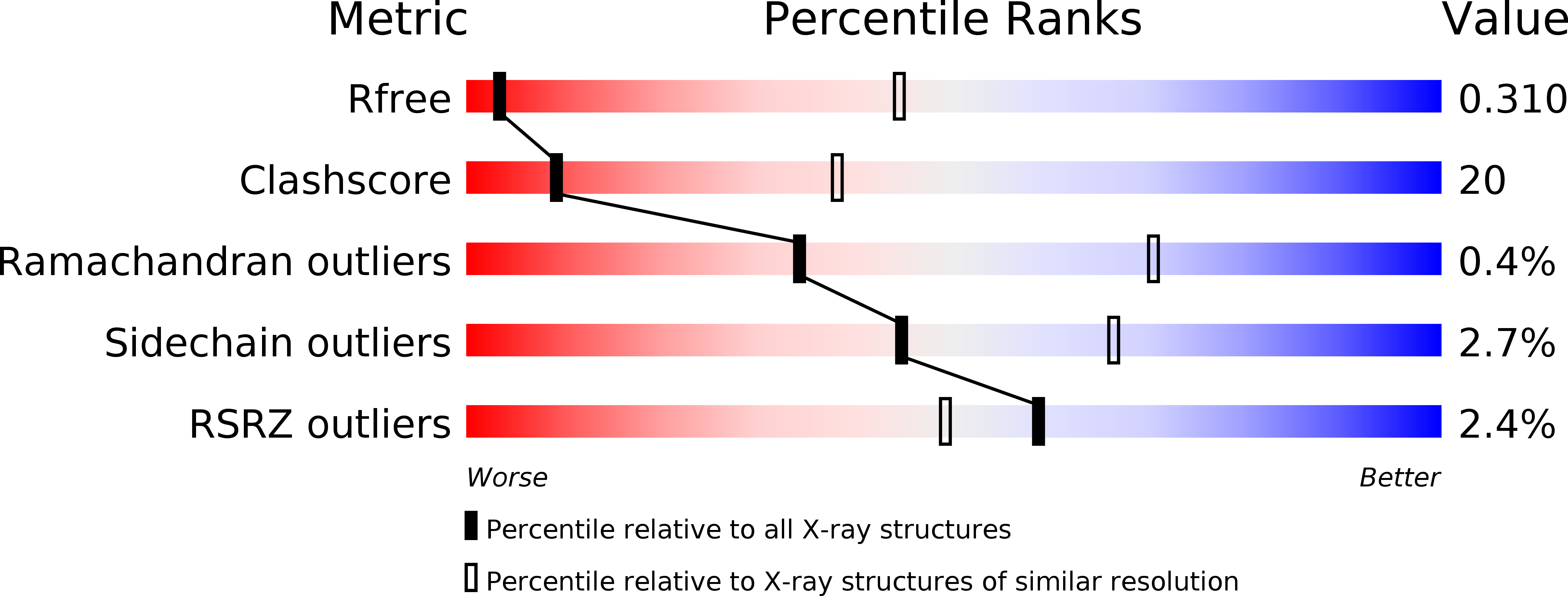
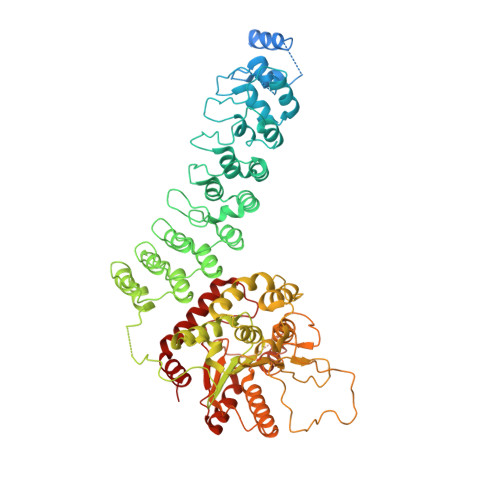
The structure of iPLA2beta reveals dimeric active sites and suggests mechanisms of regulation and localization.
Malley, K.R., Koroleva, O., Miller, I., Sanishvili, R., Jenkins, C.M., Gross, R.W., Korolev, S.(2018) Nat Commun 9: 765-765
- PubMed: 29472584
- DOI: https://doi.org/10.1038/s41467-018-03193-0
- Primary Citation of Related Structures:
6AUN - PubMed Abstract:
Calcium-independent phospholipase A 2 β (iPLA 2 β) regulates important physiological processes including inflammation, calcium homeostasis and apoptosis. It is genetically linked to neurodegenerative disorders including Parkinson's disease. Despite its known enzymatic activity, the mechanisms underlying iPLA 2 β-induced pathologic phenotypes remain poorly understood. Here, we present a crystal structure of iPLA 2 β that significantly revises existing mechanistic models. The catalytic domains form a tight dimer. They are surrounded by ankyrin repeat domains that adopt an outwardly flared orientation, poised to interact with membrane proteins. The closely integrated active sites are positioned for cooperative activation and internal transacylation. The structure and additional solution studies suggest that both catalytic domains can be bound and allosterically inhibited by a single calmodulin. These features suggest mechanisms of iPLA 2 β cellular localization and activity regulation, providing a basis for inhibitor development. Furthermore, the structure provides a framework to investigate the role of neurodegenerative mutations and the function of iPLA 2 β in the brain.
Organizational Affiliation:
Edward A. Doisy Department of Biochemistry and Molecular Biology, Saint Louis University School of Medicine, St. Louis, MO, 63104, USA.