The crystal structure of 3CL MainPro of SARS-CoV-2 with C145S mutation
Tan, K., Maltseva, N.I., Welk, L.F., Jedrzejczak, R.P., Joachimiak, A.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 3C-like proteinase | 306 | Severe acute respiratory syndrome coronavirus 2 | Mutation(s): 1 Gene Names: rep, 1a-1b EC: 3.4.22.69 | 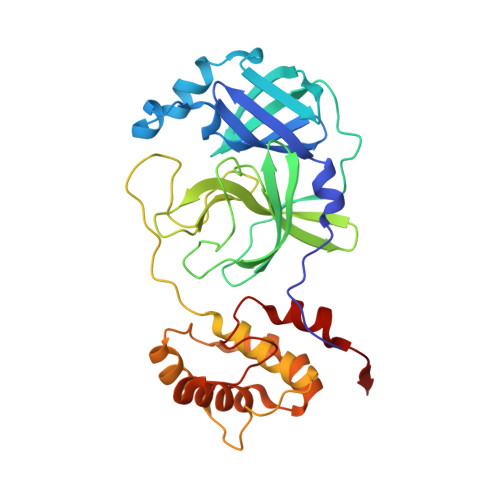 | |
UniProt | |||||
Find proteins for P0DTD1 (Severe acute respiratory syndrome coronavirus 2) Explore P0DTD1 Go to UniProtKB: P0DTD1 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0DTD1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| EDO Query on EDO | E [auth A], F [auth C], G [auth D] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 62.914 | α = 77.93 |
| b = 67.85 | β = 89.55 |
| c = 77.886 | γ = 72.9 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| SBC-Collect | data collection |
| HKL-3000 | data reduction |
| HKL-3000 | phasing |
| HKL-3000 | data scaling |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | HHSN272201 700060C |