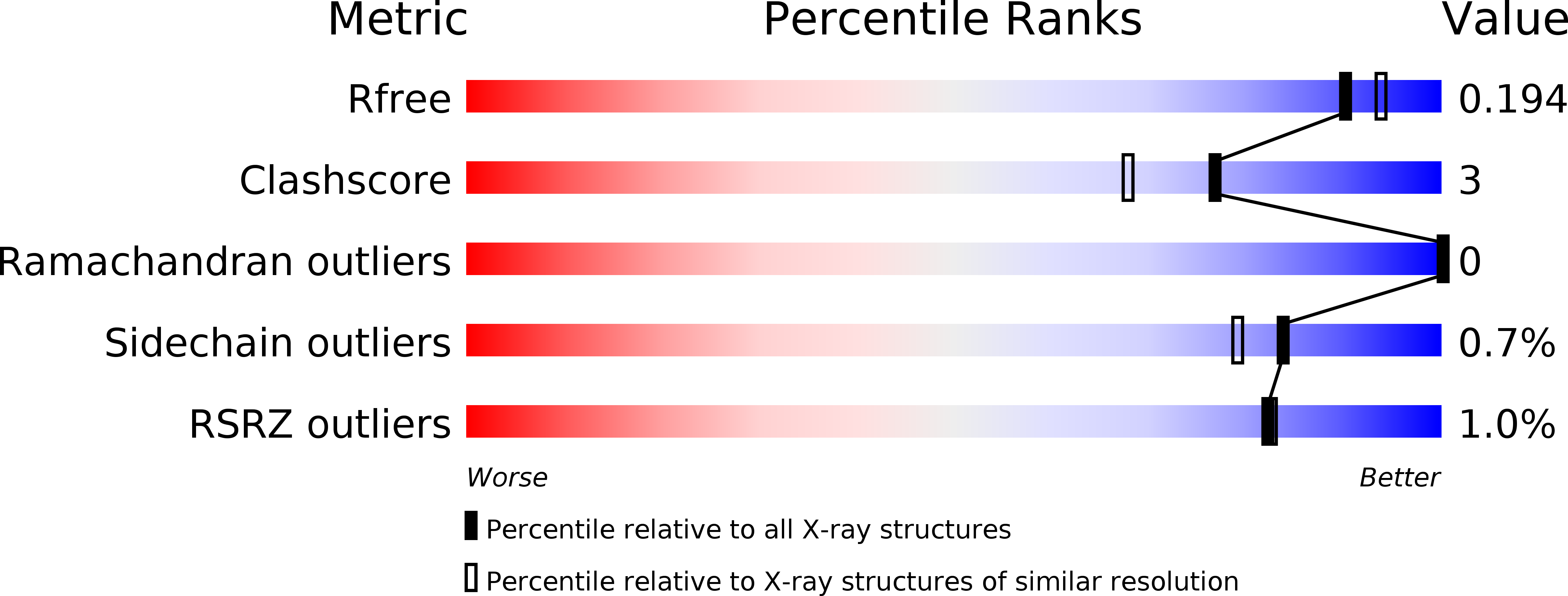
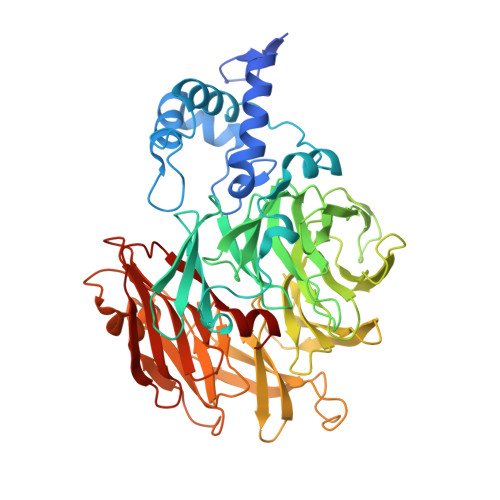
Structure of heme d 1 -free cd 1 nitrite reductase NirS.
Klunemann, T., Blankenfeldt, W.(2020) Acta Crystallogr F Struct Biol Commun 76: 250-256
- PubMed: 32510465
- DOI: https://doi.org/10.1107/S2053230X20006676
- Primary Citation of Related Structures:
6TPO, 6TSI - PubMed Abstract:
A key step in anaerobic nitrate respiration is the reduction of nitrite to nitric oxide, which is catalysed by the cd 1 nitrite reductase NirS in, for example, the Gram-negative opportunistic pathogen Pseudomonas aeruginosa. Each subunit of this homodimeric enzyme consists of a cytochrome c domain and an eight-bladed β-propeller that binds the uncommon isobacteriochlorin heme d 1 as an essential part of its active site. Although NirS has been well studied mechanistically and structurally, the focus of previous studies has been on the active heme d 1 -bound form. The heme d 1 -free form of NirS reported here, which represents a premature state of the reductase, adopts an open conformation with the cytochrome c domains moved away from each other with respect to the active enzyme. Further, the movement of a loop around Trp498 seems to be related to a widening of the propeller, allowing easier access to the heme d 1 -binding side. Finally, a possible link between the open conformation of NirS and flagella formation in P. aeruginosa is discussed.
Organizational Affiliation:
Structure and Function of Proteins, Helmholtz Centre for Infection Research, Inhoffenstrasse 7, 38124 Braunschweig, Niedersachsen, Germany.