An Asymmetric Reductase That Intercepts Acyclic Imino Acids Producedin Situby a Partner Oxidase.
Guo, J., Higgins, M.A., Daniel-Ivad, P., Ryan, K.S.(2019) J Am Chem Soc 141: 12258-12267
- PubMed: 31298853
- DOI: https://doi.org/10.1021/jacs.9b03307
- Primary Citation of Related Structures:
6P2I - PubMed Abstract:
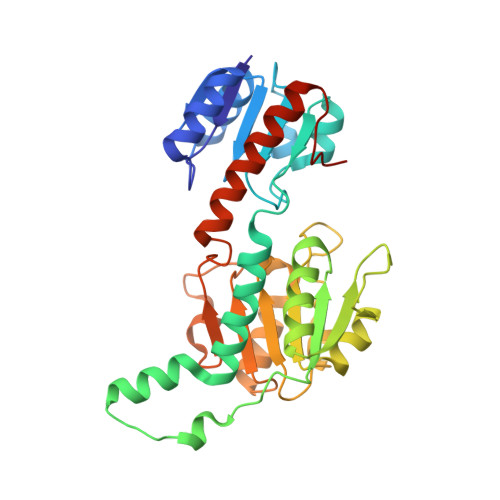
Acyclic imines are unstable in aqueous conditions. For this reason, known imine reductases, which enable the synthesis of chiral amines, mainly intercept stable cyclic imines. Here we report the detailed biochemical and structural characterization of Bsp5, an imino acid reductase from the d-2-hydroxyacid dehydrogenase family that reduces acyclic imino acids produced in situ by a partner oxidase. We determine a 1.6 Å resolution structure of Bsp5 in complex with d-arginine and coenzyme NADPH. Combined with mutagenesis work, our study reveals the minimal structural constraints for its biosynthetic activity. Furthermore, we demonstrate that Bsp5 can intercept more complex products from an alternate oxidase partner, suggesting that this oxidase-imino acid reductase pair could be evolved for biocatalytic conversion of l-amino acids to d-amino acids.
Organizational Affiliation:
Department of Chemistry , University of British Columbia , Vancouver , British Columbia V6T 1Z4 , Canada.