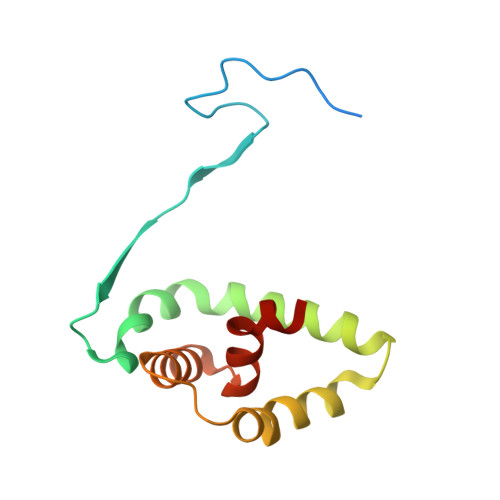
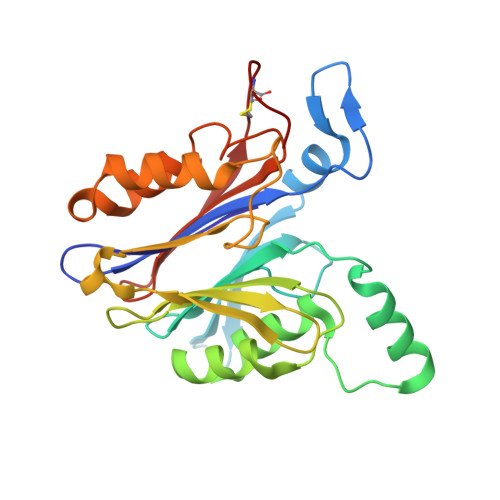
Molecular Mechanism of Inhibition of Acid Ceramidase by Carmofur.
Dementiev, A., Joachimiak, A., Nguyen, H., Gorelik, A., Illes, K., Shabani, S., Gelsomino, M., Ahn, E.E., Nagar, B., Doan, N.(2019) J Med Chem 62: 987-992
- PubMed: 30525581
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01723
- Primary Citation of Related Structures:
6MHM - PubMed Abstract:
Human acid ceramidase (AC) is a lysosomal cysteine amidase, which has received a great deal of interest in recent years as a potential target for the development of new therapeutics against melanoma and glioblastoma tumors. Despite the strong interest in obtaining structural information, only the structures of the apo-AC enzyme in its zymogen and activated conformations are available. In this work, the crystal structure of AC in complex with the covalent carmofur inhibitor is presented. Carmofur is an antineoplastic drug containing an electrophilic carbonyl reactive group that targets the catalytic cysteine. This novel structural data explains the basis of the AC inhibition, provides insights into the enzymatic properties of the protein, and is a great aid toward the structure-based drug design of potent inhibitors for AC, providing the detailed mechanism, which has eluded the scientific community for more than 30 years, of carmofur's mysterious 5-fluorouracil-independent antitumor activity.
Organizational Affiliation:
Structural Biology Center, Biosciences Division , Argonne National Laboratory , Lemont , Illinois 60439 , United States.