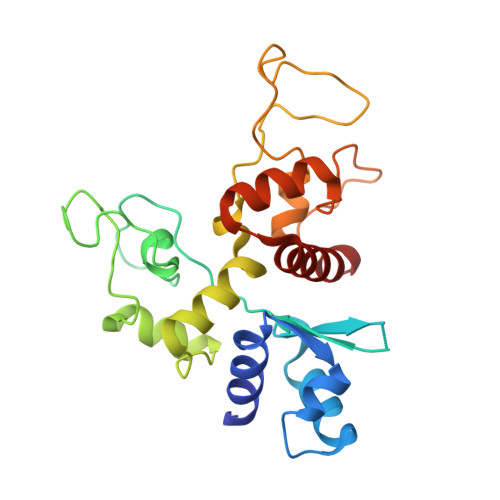
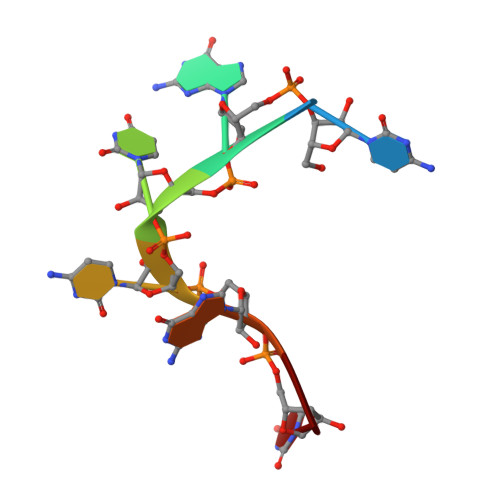
Molecular Mechanism of RNA Recognition by Zinc-Finger Antiviral Protein.
Luo, X., Wang, X., Gao, Y., Zhu, J., Liu, S., Gao, G., Gao, P.(2020) Cell Rep 30: 46-52.e4
- PubMed: 31914396
- DOI: https://doi.org/10.1016/j.celrep.2019.11.116
- Primary Citation of Related Structures:
6L1W - PubMed Abstract:
Zinc-finger antiviral protein (ZAP) is a host antiviral factor that specifically restricts a wide range of viruses. ZAP selectively binds to CG-dinucleotide-enriched RNA sequences and recruits multiple RNA degradation machines to degrade target viral RNA. However, the molecular mechanism and structural basis for ZAP recognition of specific RNA are not clear. Here, we report the crystal structure of the ZAP N-terminal domain bound to a CG-rich single-stranded RNA, providing the molecular basis for its specific recognition of a CG dinucleotide and additional guanine and cytosine. The four zinc fingers of ZAP adopt a unique architecture and form extensive interactions with RNA. Mutations of both protein and RNA at the RNA-ZAP interacting surface reduce the in vitro binding affinity and cellular antiviral activity. This work reveals the molecular mechanism of ZAP recognition of specific target RNA and also provides insights into the mechanism by which ZAP coordinates downstream RNA degradation processes.
Organizational Affiliation:
Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230026, China; CAS Key Laboratory of Infection and Immunity, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China; National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China.