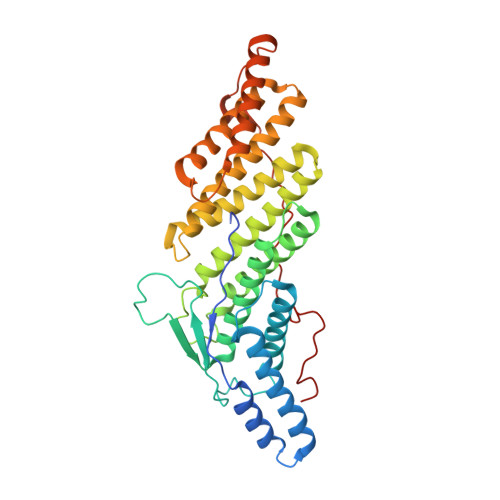
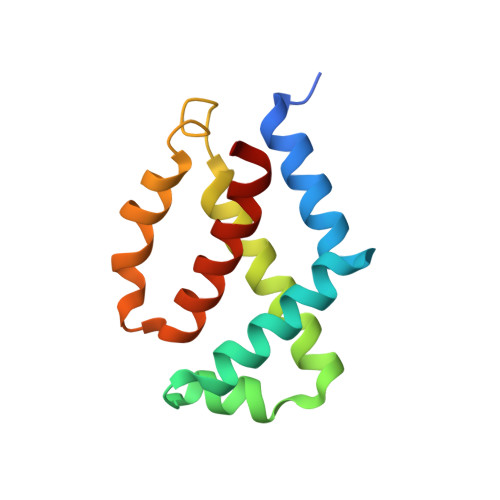
Structural Insight into the Interaction of Sendai Virus C Protein with Alix To Stimulate Viral Budding.
Oda, K., Matoba, Y., Sugiyama, M., Sakaguchi, T.(2021) J Virol : JVI0081521-JVI0081521
- PubMed: 34287046
- DOI: https://doi.org/10.1128/JVI.00815-21
- Primary Citation of Related Structures:
6KP3 - PubMed Abstract:
Sendai virus (SeV), belonging to the Respirovirus genus of the family Paramyxoviridae , harbors an accessory protein, named C protein, which facilitates viral pathogenicity in mice. In addition, the C protein is known to stimulate the budding of virus-like particles by binding to the host ALG-2 interacting protein X (Alix), a component of the endosomal sorting complexes required for transport (ESCRT) machinery. However, small interfering RNA (siRNA)-mediated gene knockdown studies suggested that neither Alix nor C protein is related to SeV budding. In the present study, we determined the crystal structure of a complex comprising the C-terminal half of the C protein (Y3) and the Bro1 domain of Alix at a resolution of 2.2 Å to investigate the role of the complex in SeV budding. The structure revealed that a novel consensus sequence, LXXW, which is conserved among Respirovirus C proteins, is important for Alix binding. SeV possessing a mutated C protein with reduced Alix-binding affinity showed impaired virus production, which correlated with the binding affinity. Infectivity analysis showed a 160-fold reduction at 12 h postinfection compared with nonmutated virus, while C protein competes with CHMP4, one subunit of the ESCRT-III complex, for binding to Alix. All together, these results highlight the critical role of C protein in SeV budding. IMPORTANCE Human parainfluenza virus type I (hPIV1) is a respiratory pathogen affecting young children, immunocompromised patients, and the elderly, with no available vaccines or antiviral drugs. Sendai virus (SeV), a murine counterpart of hPIV1, has been studied extensively to determine the molecular and biological properties of hPIV1. These viruses possess a multifunctional accessory protein, C protein, which is essential for stimulating viral reproduction, but its role in budding remains controversial. In the present study, the crystal structure of the C-terminal half of the SeV C protein associated with the Bro1 domain of Alix, a component of cell membrane modulating machinery ESCRT, was elucidated. Based on the structure, we designed mutant C proteins with different binding affinities to Alix and showed that the interaction between C and Alix is vital for viral budding. These findings provide new insights into the development of new antiviral drugs against hPIV1.
Organizational Affiliation:
Department of Virology, Institute of Biomedical & Health Sciences, Hiroshima University, Hiroshima, Japan.