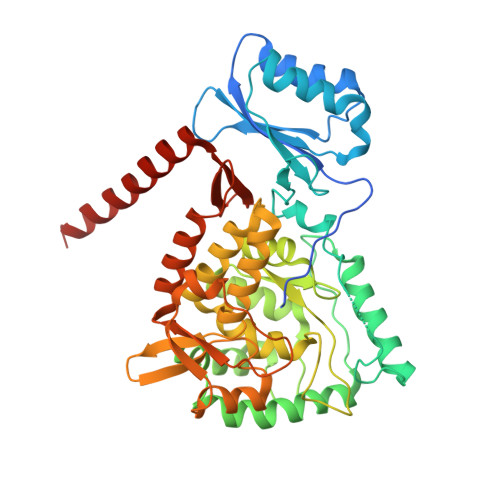
Structure of full-length human phenylalanine hydroxylase in complex with tetrahydrobiopterin.
Flydal, M.I., Alcorlo-Pages, M., Johannessen, F.G., Martinez-Caballero, S., Skjærven, L., Fernandez-Leiro, R., Martinez, A., Hermoso, J.A.(2019) Proc Natl Acad Sci U S A 116: 11229-11234
- PubMed: 31118288
- DOI: https://doi.org/10.1073/pnas.1902639116
- Primary Citation of Related Structures:
6HPO, 6HYC - PubMed Abstract:
Phenylalanine hydroxylase (PAH) is a key enzyme in the catabolism of phenylalanine, and mutations in this enzyme cause phenylketonuria (PKU), a genetic disorder that leads to brain damage and mental retardation if untreated. Some patients benefit from supplementation with a synthetic formulation of the cofactor tetrahydrobiopterin (BH 4 ) that partly acts as a pharmacological chaperone. Here we present structures of full-length human PAH (hPAH) both unbound and complexed with BH 4 in the precatalytic state. Crystal structures, solved at 3.18-Å resolution, show the interactions between the cofactor and PAH, explaining the negative regulation exerted by BH 4 BH 4 forms several H-bonds with the N-terminal autoregulatory tail but is far from the catalytic Fe II Upon BH 4 binding a polar and salt-bridge interaction network links the three PAH domains, explaining the stability conferred by BH 4 Importantly, BH 4 binding modulates the interaction between subunits, providing information about PAH allostery. Moreover, we also show that the cryo-EM structure of hPAH in absence of BH 4 reveals a highly dynamic conformation for the tetramers. Structural analyses of the hPAH:BH 4 subunits revealed that the substrate-induced movement of Tyr138 into the active site could be coupled to the displacement of BH 4 from the precatalytic toward the active conformation, a molecular mechanism that was supported by site-directed mutagenesis and targeted molecular dynamics simulations. Finally, comparison of the rat and human PAH structures show that hPAH is more dynamic, which is related to amino acid substitutions that enhance the flexibility of hPAH and may increase the susceptibility to PKU-associated mutations.
Organizational Affiliation:
Department of Biomedicine, University of Bergen, 5009 Bergen, Norway.