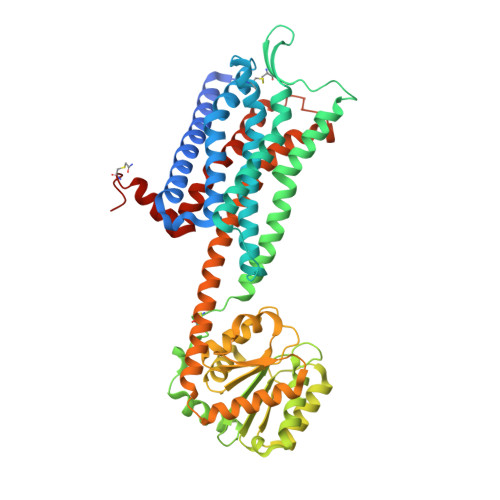
Crystal structures of the human neurokinin 1 receptor in complex with clinically used antagonists.
Schoppe, J., Ehrenmann, J., Klenk, C., Rucktooa, P., Schutz, M., Dore, A.S., Pluckthun, A.(2019) Nat Commun 10: 17-17
- PubMed: 30604743
- DOI: https://doi.org/10.1038/s41467-018-07939-8
- Primary Citation of Related Structures:
6HLL, 6HLO, 6HLP - PubMed Abstract:
Neurokinins (or tachykinins) are peptides that modulate a wide variety of human physiology through the neurokinin G protein-coupled receptor family, implicated in a diverse array of pathological processes. Here we report high-resolution crystal structures of the human NK 1 receptor (NK 1 R) bound to two small-molecule antagonist therapeutics - aprepitant and netupitant and the progenitor antagonist CP-99,994. The structures reveal the detailed interactions between clinically approved antagonists and NK 1 R, which induce a distinct receptor conformation resulting in an interhelical hydrogen-bond network that cross-links the extracellular ends of helices V and VI. Furthermore, the high-resolution details of NK 1 R bound to netupitant establish a structural rationale for the lack of basal activity in NK 1 R. Taken together, these co-structures provide a comprehensive structural basis of NK 1 R antagonism and will facilitate the design of new therapeutics targeting the neurokinin receptor family.
Organizational Affiliation:
Department of Biochemistry, University of Zürich, Winterthurerstrasse 190, CH-8057, Zürich, Switzerland.