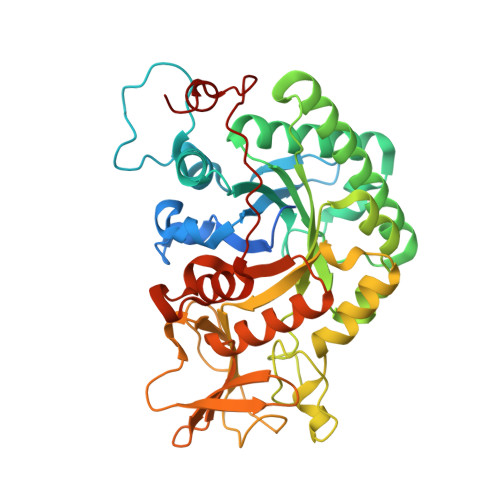
Use of chitin and chitosan to produce new chitooligosaccharides by chitinase Chit42: enzymatic activity and structural basis of protein specificity.
Kidibule, P.E., Santos-Moriano, P., Jimenez-Ortega, E., Ramirez-Escudero, M., Limon, M.C., Remacha, M., Plou, F.J., Sanz-Aparicio, J., Fernandez-Lobato, M.(2018) Microb Cell Fact 17: 47-47
- PubMed: 29566690
- DOI: https://doi.org/10.1186/s12934-018-0895-x
- Primary Citation of Related Structures:
6EPB - PubMed Abstract:
Chitinases are ubiquitous enzymes that have gained a recent biotechnological attention due to their ability to transform biological waste from chitin into valued chito-oligomers with wide agricultural, industrial or medical applications. The biological activity of these molecules is related to their size and acetylation degree. Chitinase Chit42 from Trichoderma harzianum hydrolyses chitin oligomers with a minimal of three N-acetyl-D-glucosamine (GlcNAc) units. Gene chit42 was previously characterized, and according to its sequence, the encoded protein included in the structural Glycoside Hydrolase family GH18.
Organizational Affiliation:
Department of Molecular Biology, Centre for Molecular Biology Severo Ochoa (CSIC-UAM), University Autonomous from Madrid, C/ Nicolás Cabrera, 1, Cantoblanco, 28049, Madrid, Spain.