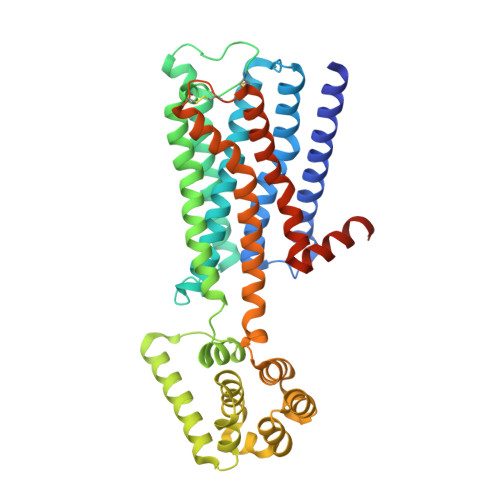
Structure-guided development of selective M3 muscarinic acetylcholine receptor antagonists
Liu, H., Hofmann, J., Fish, I., Schaake, B., Eitel, K., Bartuschat, A., Kaindl, J., Rampp, H., Banerjee, A., Hubner, H., Clark, M.J., Vincent, S.G., Fisher, J.T., Heinrich, M.R., Hirata, K., Liu, X., Sunahara, R.K., Shoichet, B.K., Kobilka, B.K., Gmeiner, P.(2018) Proc Natl Acad Sci U S A 115: 12046-12050
- PubMed: 30404914
- DOI: https://doi.org/10.1073/pnas.1813988115
- Primary Citation of Related Structures:
5ZHP - PubMed Abstract:
Drugs that treat chronic obstructive pulmonary disease by antagonizing the M3 muscarinic acetylcholine receptor (M3R) have had a significant effect on health, but can suffer from their lack of selectivity against the M2R subtype, which modulates heart rate. Beginning with the crystal structures of M2R and M3R, we exploited a single amino acid difference in their orthosteric binding pockets using molecular docking and structure-based design. The resulting M3R antagonists had up to 100-fold selectivity over M2R in affinity and over 1,000-fold selectivity in vivo. The crystal structure of the M3R-selective antagonist in complex with M3R corresponded closely to the docking-predicted geometry, providing a template for further optimization.
Organizational Affiliation:
Beijing Advanced Innovation Center for Structural Biology, School of Medicine, Tsinghua University, 100084 Beijing, China.