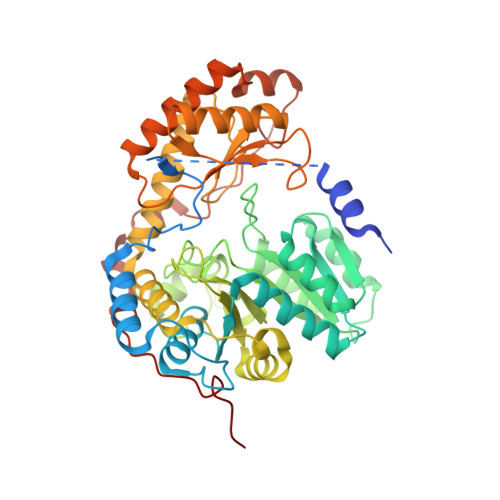
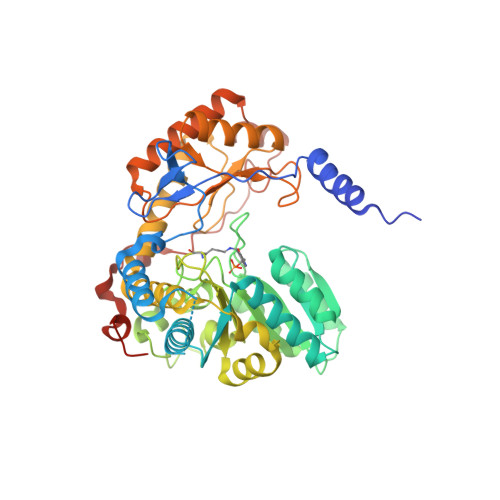
Structure of the Mitochondrial Aminolevulinic Acid Synthase, a Key Heme Biosynthetic Enzyme.
Brown, B.L., Kardon, J.R., Sauer, R.T., Baker, T.A.(2018) Structure 26: 580-589.e4
- PubMed: 29551290
- DOI: https://doi.org/10.1016/j.str.2018.02.012
- Primary Citation of Related Structures:
5TXR, 5TXT - PubMed Abstract:
5-Aminolevulinic acid synthase (ALAS) catalyzes the first step in heme biosynthesis. We present the crystal structure of a eukaryotic ALAS from Saccharomyces cerevisiae. In this homodimeric structure, one ALAS subunit contains covalently bound cofactor, pyridoxal 5'-phosphate (PLP), whereas the second is PLP free. Comparison between the subunits reveals PLP-coupled reordering of the active site and of additional regions to achieve the active conformation of the enzyme. The eukaryotic C-terminal extension, a region altered in multiple human disease alleles, wraps around the dimer and contacts active-site-proximal residues. Mutational analysis demonstrates that this C-terminal region that engages the active site is important for ALAS activity. Our discovery of structural elements that change conformation upon PLP binding and of direct contact between the C-terminal extension and the active site thus provides a structural basis for investigation of disruptions in the first step of heme biosynthesis and resulting human disorders.
Organizational Affiliation:
Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 02139, USA.