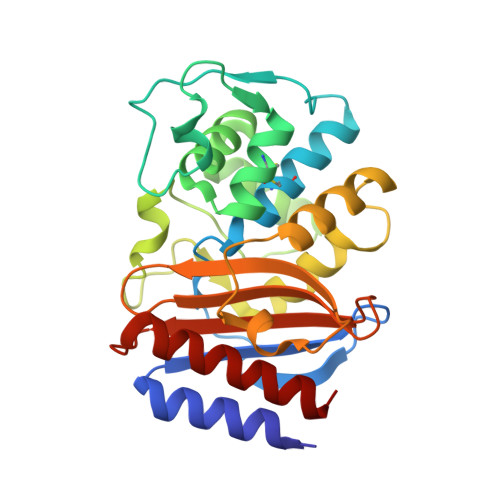
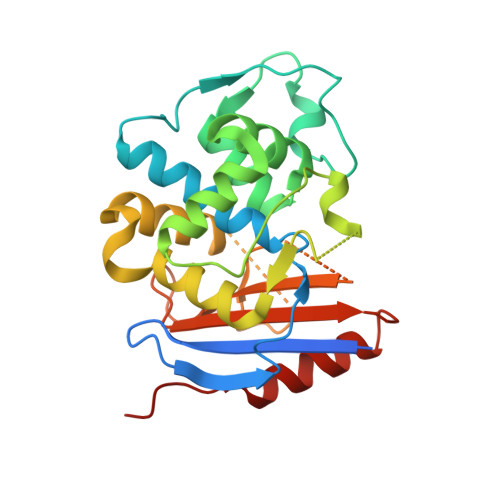
Promiscuous Protein Binding as a Function of Protein Stability.
Cohen-Khait, R., Dym, O., Hamer-Rogotner, S., Schreiber, G.(2017) Structure 25: 1867-1874.e3
- PubMed: 29211984
- DOI: https://doi.org/10.1016/j.str.2017.11.002
- Primary Citation of Related Structures:
5NPO - PubMed Abstract:
Proteins have evolved to balance efficient binding of desired partners with rejection of unwanted interactions. To investigate the evolution of protein-protein interactions, we selected a random library of pre-stabilized TEM1 β-lactamase against wild-type TEM1 using yeast surface display. Three mutations were sufficient to achieve micromolar affinity binding between the two. The X-ray structure emphasized that the main contribution of the selected mutations was to modify the protein fold, specifically removing the N'-terminal helix, which consequently allowed protein coupling via a β-sheet-mediated interaction resembling amyloid interaction mode. The only selected mutation located at the interaction interface (E58V) is reminiscent of the single mutation commonly causing sickle-cell anemia. Interestingly, the evolved mutations cannot be inserted into the wild-type protein due to reduced thermal stability of the resulting mutant protein. These results reveal a simple mechanism by which undesirable binding is purged by loss of thermal stability.
Organizational Affiliation:
Department of Biomolecular Sciences, Weizmann Institute of Science, 76100 Rehovot, Israel.