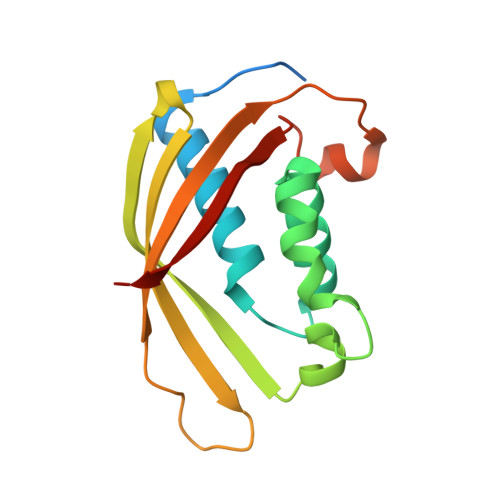
Structural Analysis and Inhibition of TraE from the pKM101 Type IV Secretion System.
Casu, B., Smart, J., Hancock, M.A., Smith, M., Sygusch, J., Baron, C.(2016) J Biol Chem 291: 23817-23829
- PubMed: 27634044
- DOI: https://doi.org/10.1074/jbc.M116.753327
- Primary Citation of Related Structures:
5I97 - PubMed Abstract:
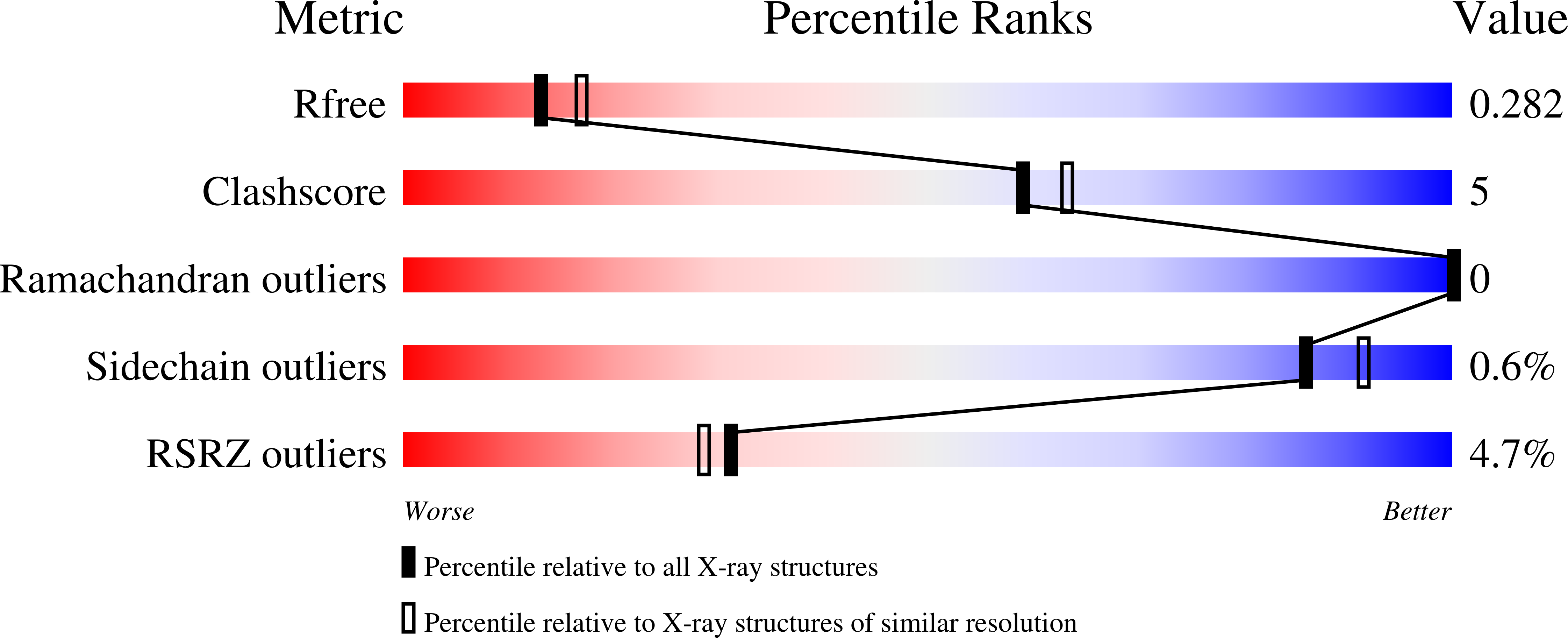
Gram-negative bacteria use type IV secretion systems (T4SSs) for a variety of macromolecular transport processes that include the exchange of genetic material. The pKM101 plasmid encodes a T4SS similar to the well-studied model systems from Agrobacterium tumefaciens and Brucella suis Here, we studied the structure and function of TraE, a homolog of VirB8 that is an essential component of all T4SSs. Analysis by X-ray crystallography revealed a structure that is similar to other VirB8 homologs but displayed an altered dimerization interface. The dimerization interface observed in the X-ray structure was corroborated using the bacterial two-hybrid assay, biochemical characterization of the purified protein, and in vivo complementation, demonstrating that there are different modes of dimerization among VirB8 homologs. Analysis of interactions using the bacterial two-hybrid and cross-linking assays showed that TraE and its homologs from Agrobacterium, Brucella, and Helicobacter pylori form heterodimers. They also interact with heterologous VirB10 proteins, indicating a significant degree of plasticity in the protein-protein interactions of VirB8-like proteins. To further assess common features of VirB8-like proteins, we tested a series of small molecules derived from inhibitors of Brucella VirB8 dimerization. These molecules bound to TraE in vitro, docking predicted that they bind to a structurally conserved surface groove of the protein, and some of them inhibited pKM101 plasmid transfer. VirB8-like proteins thus share functionally important sites, and these can be exploited for the design of specific inhibitors of T4SS function.
Organizational Affiliation:
From the Department of Biochemistry and Molecular Medicine, Faculty of Medicine, Université de Montréal, Montréal, Quebec H3C 3J7, Canada, and.