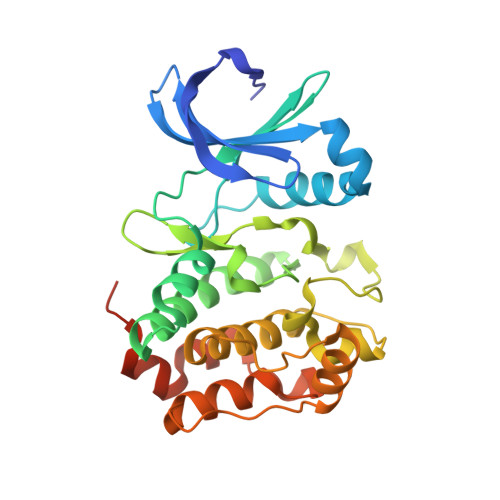
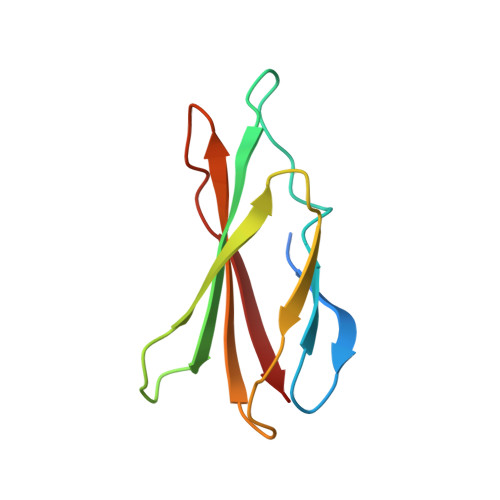
Allosteric modulation of a human protein kinase with monobodies.
Zorba, A., Nguyen, V., Koide, A., Hoemberger, M., Zheng, Y., Kutter, S., Kim, C., Koide, S., Kern, D.(2019) Proc Natl Acad Sci U S A 116: 13937-13942
- PubMed: 31239342
- DOI: https://doi.org/10.1073/pnas.1906024116
- Primary Citation of Related Structures:
5G15, 6C83 - PubMed Abstract:
Despite being the subject of intense effort and scrutiny, kinases have proven to be consistently challenging targets in inhibitor drug design. A key obstacle has been promiscuity and consequent adverse effects of drugs targeting the ATP binding site. Here we introduce an approach to controlling kinase activity by using monobodies that bind to the highly specific regulatory allosteric pocket of the oncoprotein Aurora A (AurA) kinase, thereby offering the potential for more specific kinase modulators. Strikingly, we identify a series of highly specific monobodies acting either as strong kinase inhibitors or activators via differential recognition of structural motifs in the allosteric pocket. X-ray crystal structures comparing AurA bound to activating vs inhibiting monobodies reveal the atomistic mechanism underlying allosteric modulation. The results reveal 3 major advantages of targeting allosteric vs orthosteric sites: extreme selectivity, ability to inhibit as well as activate, and avoidance of competing with ATP that is present at high concentrations in the cells. We envision that exploiting allosteric networks for inhibition or activation will provide a general, powerful pathway toward rational drug design.
Organizational Affiliation:
Department of Biochemistry, Brandeis University, Waltham, MA 02454.