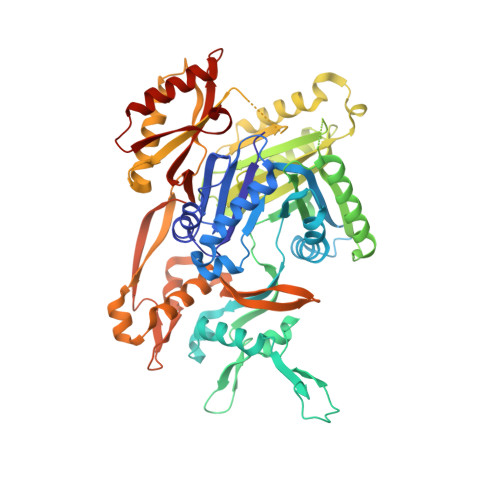
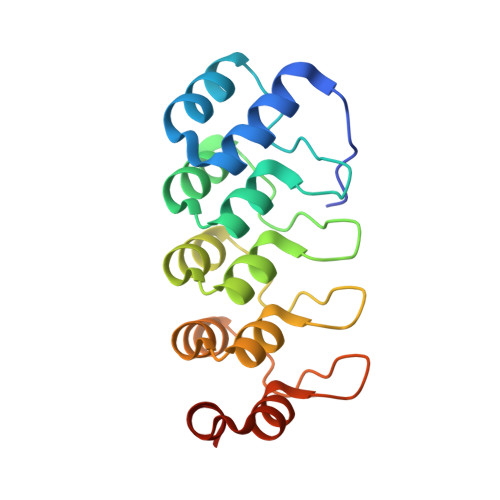
Molecular basis for inhibition of AcrB multidrug efflux pump by novel and powerful pyranopyridine derivatives.
Sjuts, H., Vargiu, A.V., Kwasny, S.M., Nguyen, S.T., Kim, H.S., Ding, X., Ornik, A.R., Ruggerone, P., Bowlin, T.L., Nikaido, H., Pos, K.M., Opperman, T.J.(2016) Proc Natl Acad Sci U S A 113: 3509-3514
- PubMed: 26976576
- DOI: https://doi.org/10.1073/pnas.1602472113
- Primary Citation of Related Structures:
5EN5, 5ENO, 5ENP, 5ENQ, 5ENR, 5ENS, 5ENT - PubMed Abstract:
The Escherichia coli AcrAB-TolC efflux pump is the archetype of the resistance nodulation cell division (RND) exporters from Gram-negative bacteria. Overexpression of RND-type efflux pumps is a major factor in multidrug resistance (MDR), which makes these pumps important antibacterial drug discovery targets. We have recently developed novel pyranopyridine-based inhibitors of AcrB, which are orders of magnitude more powerful than the previously known inhibitors. However, further development of such inhibitors has been hindered by the lack of structural information for rational drug design. Although only the soluble, periplasmic part of AcrB binds and exports the ligands, the presence of the membrane-embedded domain in AcrB and its polyspecific binding behavior have made cocrystallization with drugs challenging. To overcome this obstacle, we have engineered and produced a soluble version of AcrB [AcrB periplasmic domain (AcrBper)], which is highly congruent in structure with the periplasmic part of the full-length protein, and is capable of binding substrates and potent inhibitors. Here, we describe the molecular basis for pyranopyridine-based inhibition of AcrB using a combination of cellular, X-ray crystallographic, and molecular dynamics (MD) simulations studies. The pyranopyridines bind within a phenylalanine-rich cage that branches from the deep binding pocket of AcrB, where they form extensive hydrophobic interactions. Moreover, the increasing potency of improved inhibitors correlates with the formation of a delicate protein- and water-mediated hydrogen bond network. These detailed insights provide a molecular platform for the development of novel combinational therapies using efflux pump inhibitors for combating multidrug resistant Gram-negative pathogens.
Organizational Affiliation:
Institute of Biochemistry, Goethe University Frankfurt, D-60438 Frankfurt, Germany;