Disruption of a hydrogen bond network in human versus spider monkey cytochrome c affects heme crevice stability.
Goldes, M.E., Jeakins-Cooley, M.E., McClelland, L.J., Mou, T.C., Bowler, B.E.(2016) J Inorg Biochem 158: 62-69
- PubMed: 26775610
- DOI: https://doi.org/10.1016/j.jinorgbio.2015.12.025
- Primary Citation of Related Structures:
5DFS - PubMed Abstract:
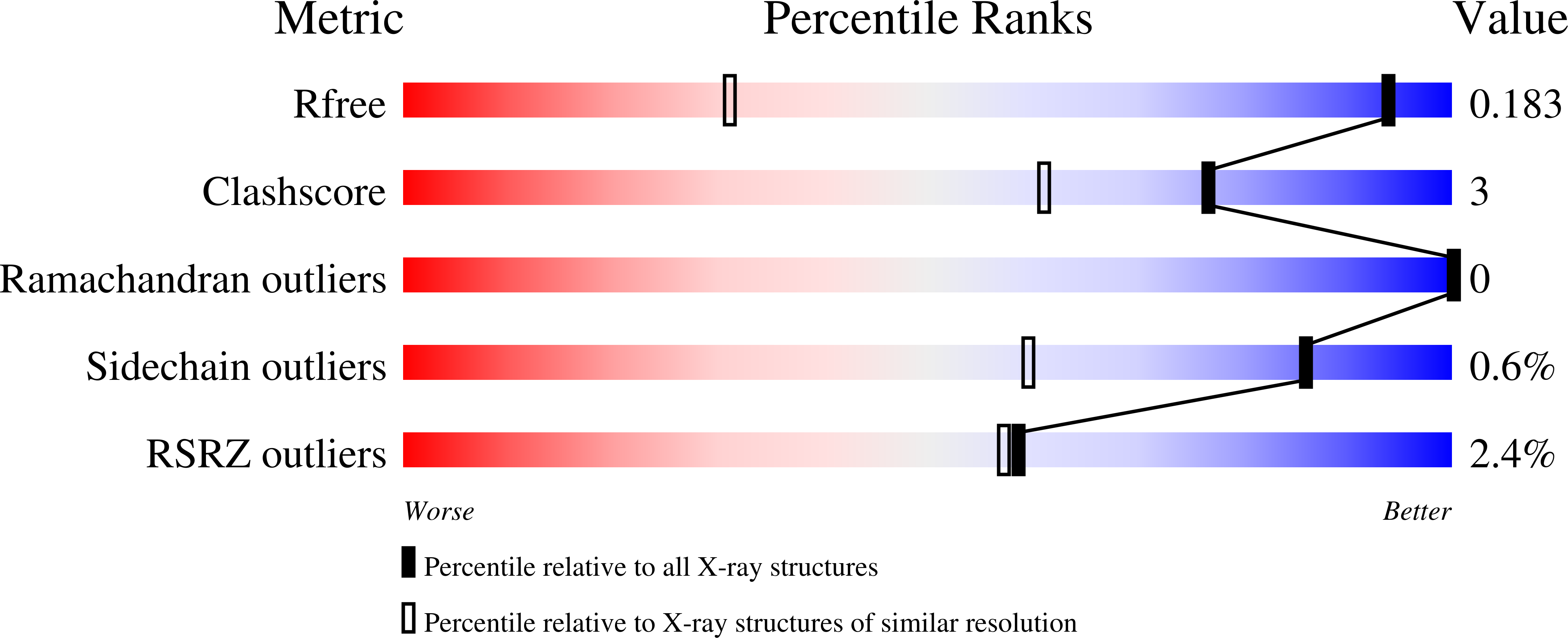
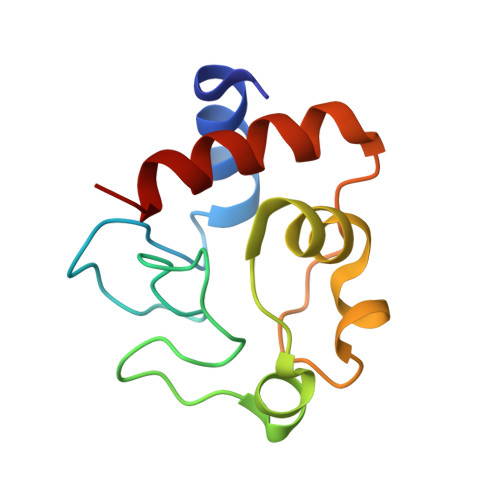
The hypothesis that the recent rapid evolution of primate cytochromes c, which primarily involves residues in the least stable Ω-loop (Ω-loop C, residues 40-57), stabilizes the heme crevice of cytochrome c relative to other mammals, is tested. To accomplish this goal, we have compared the properties of human and spider monkey cytochrome c and a set of four variants produced in the process of converting human cytochrome c into spider monkey cytochrome c. The global stability of all variants has been measured by guanidine hydrochloride denaturation. The stability of the heme crevice has been assessed with the alkaline conformational transition. Structural insight into the effects of the five amino acid substitutions needed to convert human cytochrome c into spider monkey cytochrome c is provided by a 1.15Å resolution structure of spider monkey cytochrome c. The global stability for all variants is near 9.0kcal/mol at 25°C and pH7, which is higher than that observed for other mammalian cytochromes c. The heme crevice stability is more sensitive to the substitutions required to produce spider monkey cytochrome c with decreases of up to 0.5 units in the apparent pKa of the alkaline conformational transition relative to human cytochrome c. The structure of spider monkey cytochrome c indicates that the Y46F substitution destabilizes the heme crevice by disrupting an extensive hydrogen bond network that connects three surface loops including Ω-loop D (residues 70-85), which contains the Met80 heme ligand.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Montana, Missoula, MT 59812, United States.