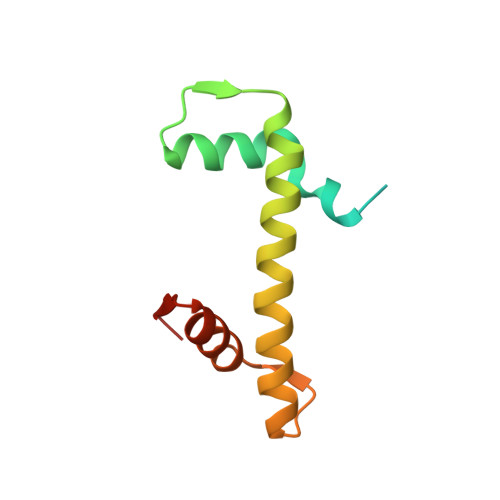
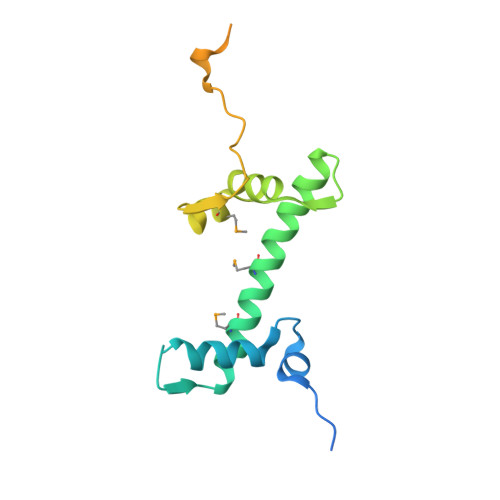
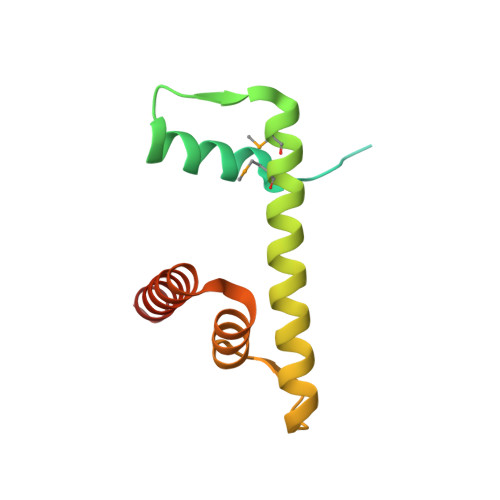
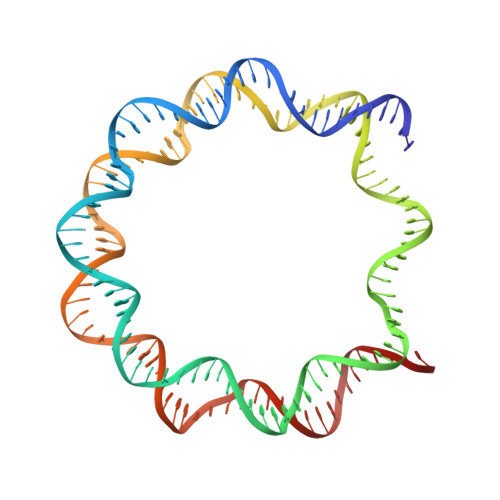
The CENP-A centromere targeting domain facilitates H4K20 monomethylation in the nucleosome by structural polymorphism.
Arimura, Y., Tachiwana, H., Takagi, H., Hori, T., Kimura, H., Fukagawa, T., Kurumizaka, H.(2019) Nat Commun 10: 576-576
- PubMed: 30718488
- DOI: https://doi.org/10.1038/s41467-019-08314-x
- Primary Citation of Related Structures:
5Z23, 5ZBX - PubMed Abstract:
Centromeric nucleosomes are composed of the centromere-specific histone H3 variant CENP-A and the core histones H2A, H2B, and H4. To establish a functional kinetochore, histone H4 lysine-20 (H4K20) must be monomethylated, but the underlying mechanism has remained enigmatic. To provide structural insights into H4K20 methylation, we here solve the crystal structure of a nucleosome containing an H3.1-CENP-A chimera, H3.1 CATD , which has a CENP-A centromere targeting domain and preserves essential CENP-A functions in vivo. Compared to the canonical H3.1 nucleosome, the H3.1 CATD nucleosome exhibits conformational changes in the H4 N-terminal tail leading to a relocation of H4K20. In particular, the H4 N-terminal tail interacts with glutamine-76 and aspartate-77 of canonical H3.1 while these interactions are cancelled in the presence of the CENP-A-specific residues valine-76 and lysine-77. Mutations of valine-76 and lysine-77 impair H4K20 monomethylation both in vitro and in vivo. These findings suggest that a CENP-A-mediated structural polymorphism may explain the preferential H4K20 monomethylation in centromeric nucleosomes.
Organizational Affiliation:
Laboratory of Chromatin Structure and Function, Institute for Quantitative Biosciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo, 113-0032, Japan.