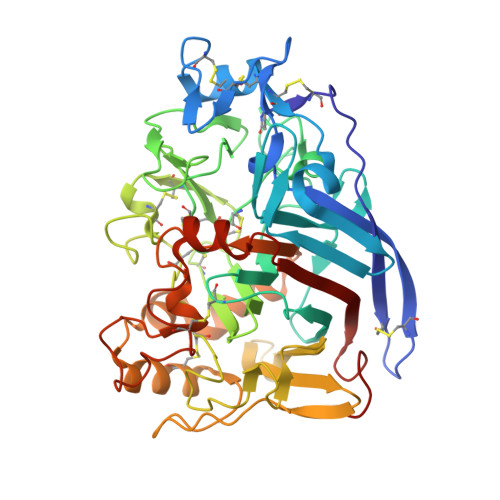
Biochemical and structural insights into a thermostable cellobiohydrolase from Myceliophthora thermophila.
Kadowaki, M.A.S., Higasi, P., de Godoy, M.O., Prade, R.A., Polikarpov, I.(2018) FEBS J 285: 559-579
- PubMed: 29222836
- DOI: https://doi.org/10.1111/febs.14356
- Primary Citation of Related Structures:
5W11 - PubMed Abstract:
Cellobiohydrolases hydrolyze cellulose, a linear polymer with glucose monomers linked exclusively by β-1,4 glycosidic linkages. The widespread hydrogen bonding network tethers individual cellulose polymers forming crystalline cellulose, which prevent the access of hydrolytic enzymes and water molecules. The most abundant enzyme secreted by Myceliophthora thermophila M77 in response to the presence of biomass is the cellobiohydrolase MtCel7A, which is composed by a GH7-catalytic domain (CD), a linker, and a CBM1-type carbohydrate-binding module. GH7 cellobiohydrolases have been studied before, and structural models have been proposed. However, currently available GH7 crystal structures only define separate catalytic domains and/or cellulose-binding modules and do not include the full-length structures that are involved in shaping the catalytic mode of operation. In this study, we determined the 3D structure of catalytic domain using X-ray crystallography and retrieved the full-length enzyme envelope via small-angle X-ray scattering (SAXS) technique. The SAXS data reveal a tadpole-like molecular shape with a rigid linker connecting the CD and CBM. Our biochemical studies show that MtCel7A has higher catalytic efficiency and thermostability as well as lower processivity when compared to the well-studied TrCel7A from Trichoderma reesei. Based on a comparison of the crystallographic structures of CDs and their molecular dynamic simulations, we demonstrate that MtCel7A has considerably higher flexibility than TrCel7A. In particular, loops that cover the active site are more flexible and undergo higher conformational fluctuations, which might account for decreased processivity and enhanced enzymatic efficiency. Our statistical coupling analysis suggests co-evolution of amino acid clusters comprising the catalytic site of MtCel7A, which correlate with the steps in the catalytic cycle of the enzyme. The atomic coordinates and structural factors of MtCel7A have been deposited in the Protein Data Bank with accession number 5W11.
Organizational Affiliation:
São Carlos Institute of Physics, University of São Paulo, Brazil.