Chemical Space Expansion of Bromodomain Ligands Guided by in Silico Virtual Couplings (AutoCouple).
Batiste, L., Unzue, A., Dolbois, A., Hassler, F., Wang, X., Deerain, N., Zhu, J., Spiliotopoulos, D., Nevado, C., Caflisch, A.(2018) ACS Cent Sci 4: 180-188
- PubMed: 29532017
- DOI: https://doi.org/10.1021/acscentsci.7b00401
- Primary Citation of Related Structures:
5NLK, 5OVB, 5OWM, 5OWW - PubMed Abstract:
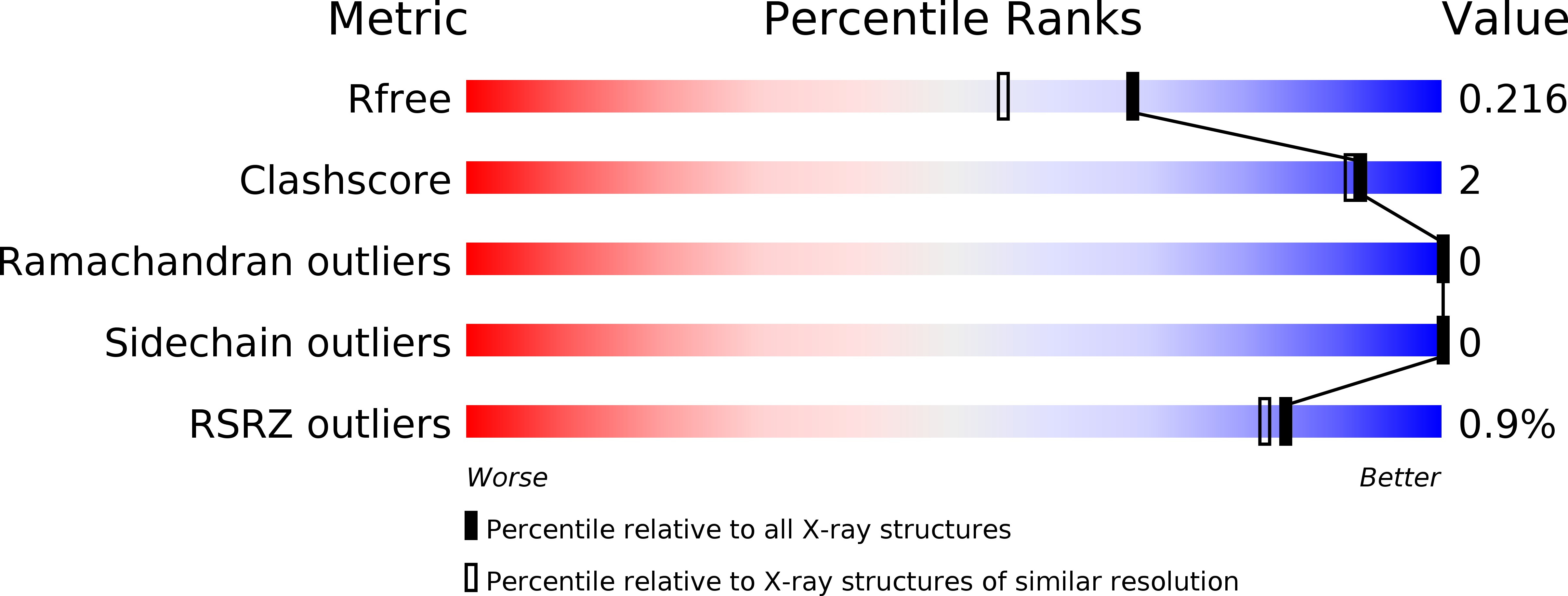
Expanding the chemical space and simultaneously ensuring synthetic accessibility is of upmost importance, not only for the discovery of effective binders for novel protein classes but, more importantly, for the development of compounds against hard-to-drug proteins. Here, we present AutoCouple, a de novo approach to computational ligand design focused on the diversity-oriented generation of chemical entities via virtual couplings. In a benchmark application, chemically diverse compounds with low-nanomolar potency for the CBP bromodomain and high selectivity against the BRD4(1) bromodomain were achieved by the synthesis of about 50 derivatives of the original fragment. The binding mode was confirmed by X-ray crystallography, target engagement in cells was demonstrated, and antiproliferative activity was showcased in three cancer cell lines. These results reveal AutoCouple as a useful in silico coupling method to expand the chemical space in hit optimization campaigns resulting in potent, selective, and cell permeable bromodomain ligands.
Organizational Affiliation:
Department of Biochemistry, University of Zurich, Winterthurerstrasse 190, CH-8057, Zürich, Switzerland.