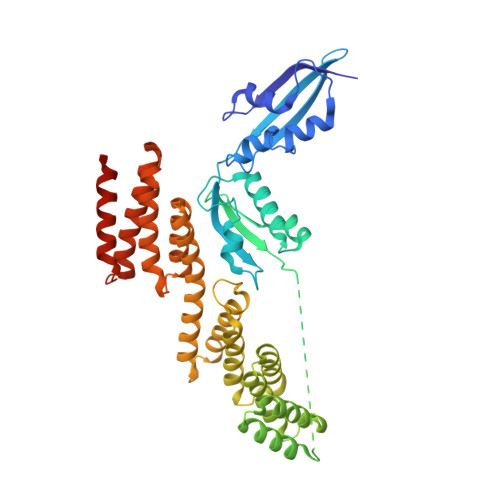
The Structure of a BamA-BamD Fusion Illuminates the Architecture of the beta-Barrel Assembly Machine Core.
Bergal, H.T., Hopkins, A.H., Metzner, S.I., Sousa, M.C.(2016) Structure 24: 243-251
- PubMed: 26749448
- DOI: https://doi.org/10.1016/j.str.2015.10.030
- Primary Citation of Related Structures:
5EFR - PubMed Abstract:
The β-barrel assembly machine (BAM) mediates folding and insertion of integral β-barrel outer membrane proteins (OMPs) in Gram-negative bacteria. Of the five BAM subunits, only BamA and BamD are essential for cell viability. Here we present the crystal structure of a fusion between BamA POTRA4-5 and BamD from Rhodothermus marinus. The POTRA5 domain binds BamD between its tetratricopeptide repeats 3 and 4. The interface structural elements are conserved in the Escherichia coli proteins, which allowed structure validation by mutagenesis and disulfide crosslinking in E. coli. Furthermore, the interface is consistent with previously reported mutations that impair BamA-BamD binding. The structure serves as a linchpin to generate a BAM model where POTRA domains and BamD form an elongated periplasmic ring adjacent to the membrane with a central cavity approximately 30 × 60 Å wide. We propose that nascent OMPs bind this periplasmic ring prior to insertion and folding by BAM.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Colorado at Boulder, Boulder, CO 80309, USA.