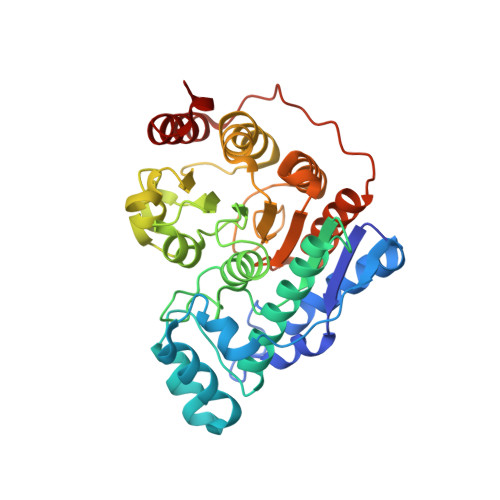
Histone deacetylase 6 structure and molecular basis of catalysis and inhibition.
Hai, Y., Christianson, D.W.(2016) Nat Chem Biol 12: 741-747
- PubMed: 27454933
- DOI: https://doi.org/10.1038/nchembio.2134
- Primary Citation of Related Structures:
5EDU, 5EEF, 5EEI, 5EEK, 5EEM, 5EEN, 5EF7, 5EF8, 5EFB, 5EFG, 5EFH, 5EFJ, 5EFK, 5EFN - PubMed Abstract:
Histone deacetylase 6 (HDAC6) is a critical target for drug design because of its role in oncogenic transformation and cancer metastasis, and is unique among all histone deacetylases in that it contains tandem catalytic domains designated CD1 and CD2. We now report the crystal structures of CD2 from Homo sapiens HDAC6 and of CD1 and CD2 from Danio rerio HDAC6. We correlated these structures with activity measurements using 13 different substrates. The catalytic activity of CD2 from both species exhibited broad substrate specificity, whereas that of CD1 was highly specific for substrates bearing C-terminal acetyllysine residues. Crystal structures of substrate complexes yielded unprecedented snapshots of the catalytic mechanism. Additionally, crystal structures of complexes with eight different inhibitors, including belinostat and panobinostat (currently used in cancer chemotherapy), the macrocyclic tetrapeptide HC toxin, and the HDAC6-specific inhibitor N-hydroxy-4-(2-((2-hydroxyethyl)(phenyl)amino)-2-oxoethyl)benzamide, revealed surprising new insight regarding changes in Zn(2+) coordination and isozyme-specific inhibition.
Organizational Affiliation:
Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, Pennsylvania, USA.