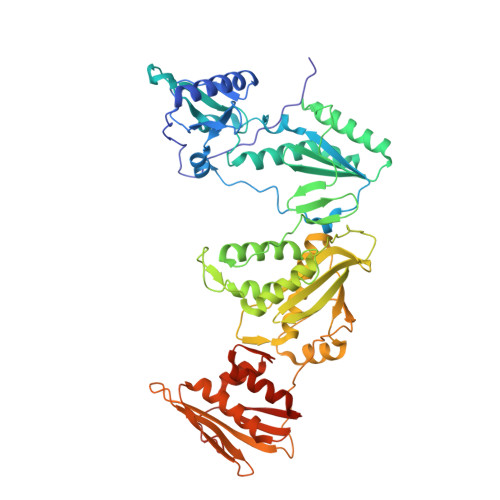
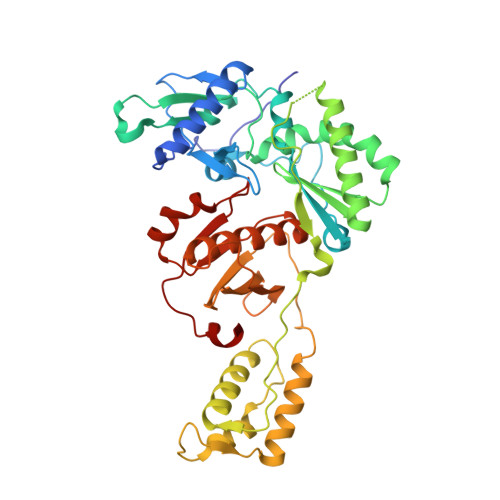
Rapid experimental SAD phasing and hot-spot identification with halogenated fragments.
Bauman, J.D., Harrison, J.J., Arnold, E.(2016) IUCrJ 3: 51-60
- PubMed: 26870381
- DOI: https://doi.org/10.1107/S2052252515021259
- Primary Citation of Related Structures:
5CXR, 5CYQ - PubMed Abstract:
Through X-ray crystallographic fragment screening, 4-bromopyrazole was discovered to be a 'magic bullet' that is capable of binding at many of the ligand 'hot spots' found in HIV-1 reverse transcriptase (RT). The binding locations can be in pockets that are 'hidden' in the unliganded crystal form, allowing rapid identification of these sites for in silico screening. In addition to hot-spot identification, this ubiquitous yet specific binding provides an avenue for X-ray crystallographic phase determination, which can be a significant bottleneck in the determination of the structures of novel proteins. The anomalous signal from 4-bromopyrazole or 4-iodopyrazole was sufficient to determine the structures of three proteins (HIV-1 RT, influenza A endonuclease and proteinase K) by single-wavelength anomalous dispersion (SAD) from single crystals. Both compounds are inexpensive, readily available, safe and very soluble in DMSO or water, allowing efficient soaking into crystals.
Organizational Affiliation:
Center for Advanced Biotechnology and Medicine, Department of Chemistry and Chemical Biology, Rutgers University , 679 Hoes Lane, Piscataway, NJ 08854, USA.