Extended Stokes shift in fluorescent proteins: chromophore-protein interactions in a near-infrared TagRFP675 variant.
Piatkevich, K.D., Malashkevich, V.N., Morozova, K.S., Nemkovich, N.A., Almo, S.C., Verkhusha, V.V.(2013) Sci Rep 3: 1847-1847
- PubMed: 23677204
- DOI: https://doi.org/10.1038/srep01847
- Primary Citation of Related Structures:
4KGE, 4KGF - PubMed Abstract:
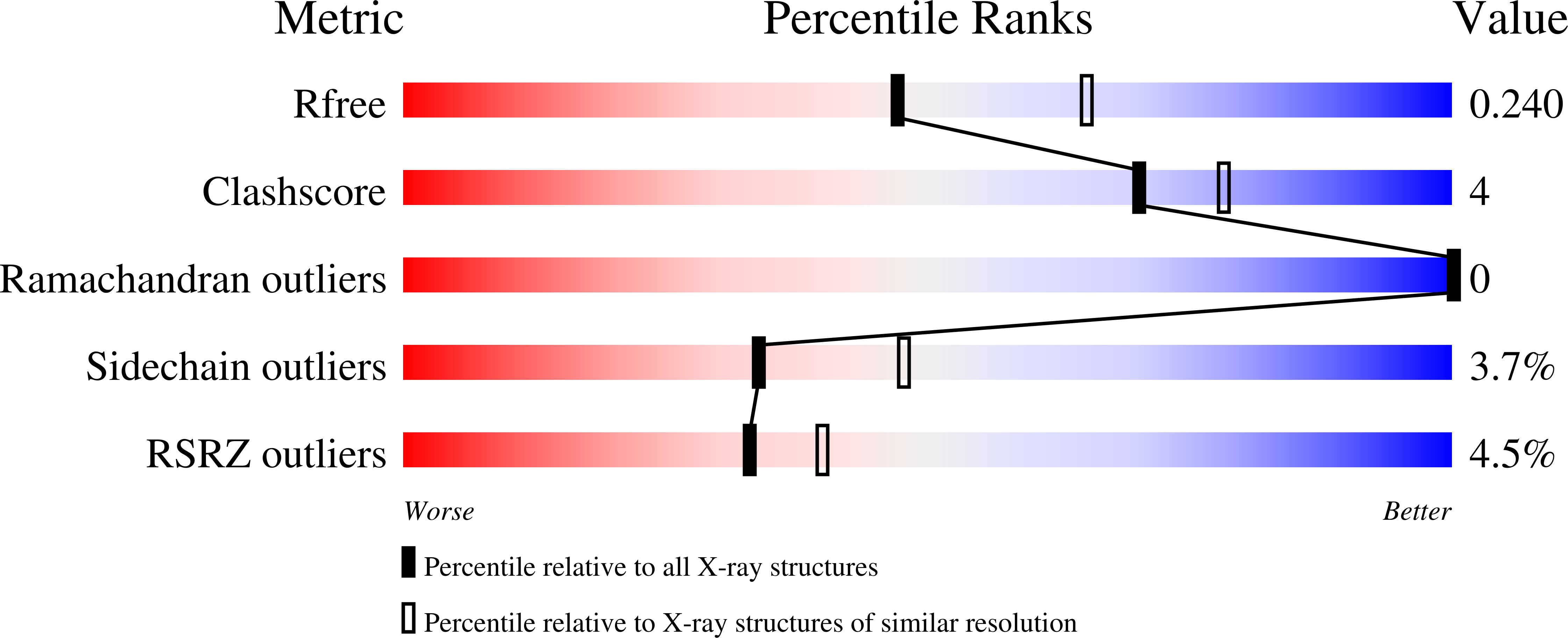
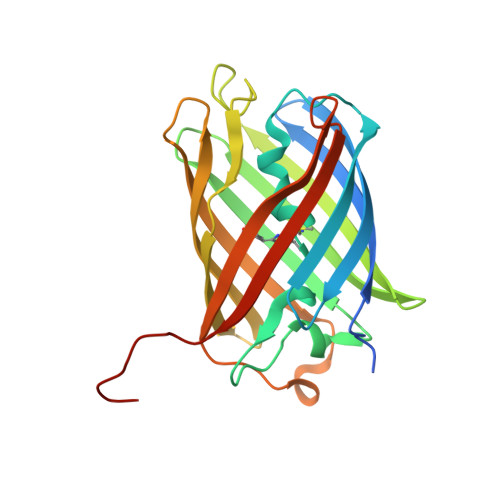
Most GFP-like fluorescent proteins exhibit small Stokes shifts (10-45 nm) due to rigidity of the chromophore environment that excludes non-fluorescent relaxation to a ground state. An unusual near-infrared derivative of the red fluorescent protein mKate, named TagRFP675, exhibits the Stokes shift, which is 30 nm extended comparing to that of the parental protein. In physiological conditions, TagRFP675 absorbs at 598 nm and emits at 675 nm that makes it the most red-shifted protein of the GFP-like protein family. In addition, its emission maximum strongly depends on the excitation wavelength. Structures of TagRFP675 revealed the common DsRed-like chromophore, which, however, interacts with the protein matrix via an extensive network of hydrogen bonds capable of large flexibility. Based on the spectroscopic, biochemical, and structural analysis we suggest that the rearrangement of the hydrogen bond interactions between the chromophore and the protein matrix is responsible for the TagRFP675 spectral properties.
Organizational Affiliation:
Department of Anatomy and Structural Biology and Gruss-Lipper Biophotonics Center, Bronx, New York 10461, United States.