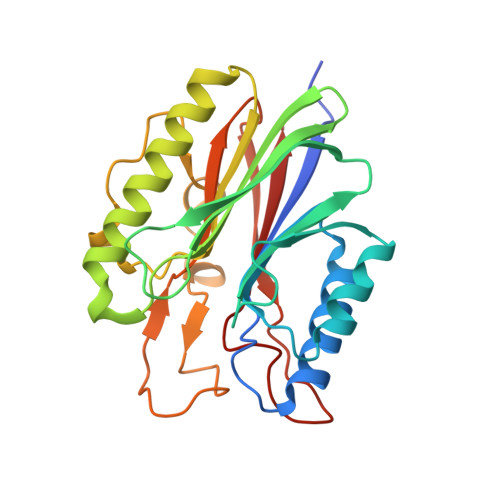
Structural basis for recognition of 5'-phosphotyrosine adducts by Tdp2.
Shi, K., Kurahashi, K., Gao, R., Tsutakawa, S.E., Tainer, J.A., Pommier, Y., Aihara, H.(2012) Nat Struct Mol Biol 19: 1372-1377
- PubMed: 23104058
- DOI: https://doi.org/10.1038/nsmb.2423
- Primary Citation of Related Structures:
4F1H, 4F1I, 4FPV, 4FVA, 4GEW - PubMed Abstract:
The DNA-repair enzyme Tdp2 resolves 5'-phosphotyrosyl DNA adducts and mediates resistance to anticancer drugs that target covalent topoisomerase-DNA complexes. Tdp2 also participates in key signaling pathways during development and tumorigenesis and cleaves a protein-RNA linkage during picornavirus replication. The crystal structure of zebrafish Tdp2 bound to DNA reveals a deep, narrow basic groove that selectively accommodates the 5' end of single-stranded DNA in a stretched conformation. The crystal structure of the full-length Caenorhabditis elegans Tdp2 shows that this groove can also accommodate an acidic peptide stretch in vitro, with glutamate and aspartate side chains occupying the DNA backbone phosphate-binding sites. This extensive molecular mimicry suggests a potential mechanism for autoregulation and interaction of Tdp2 with phosphorylated proteins in signaling. Our study provides a framework to interrogate functions of Tdp2 and develop inhibitors for chemotherapeutic and antiviral applications.
Organizational Affiliation:
Department of Biochemistry, Molecular Biology and Biophysics, University of Minnesota, Minneapolis, Minnesota, USA.