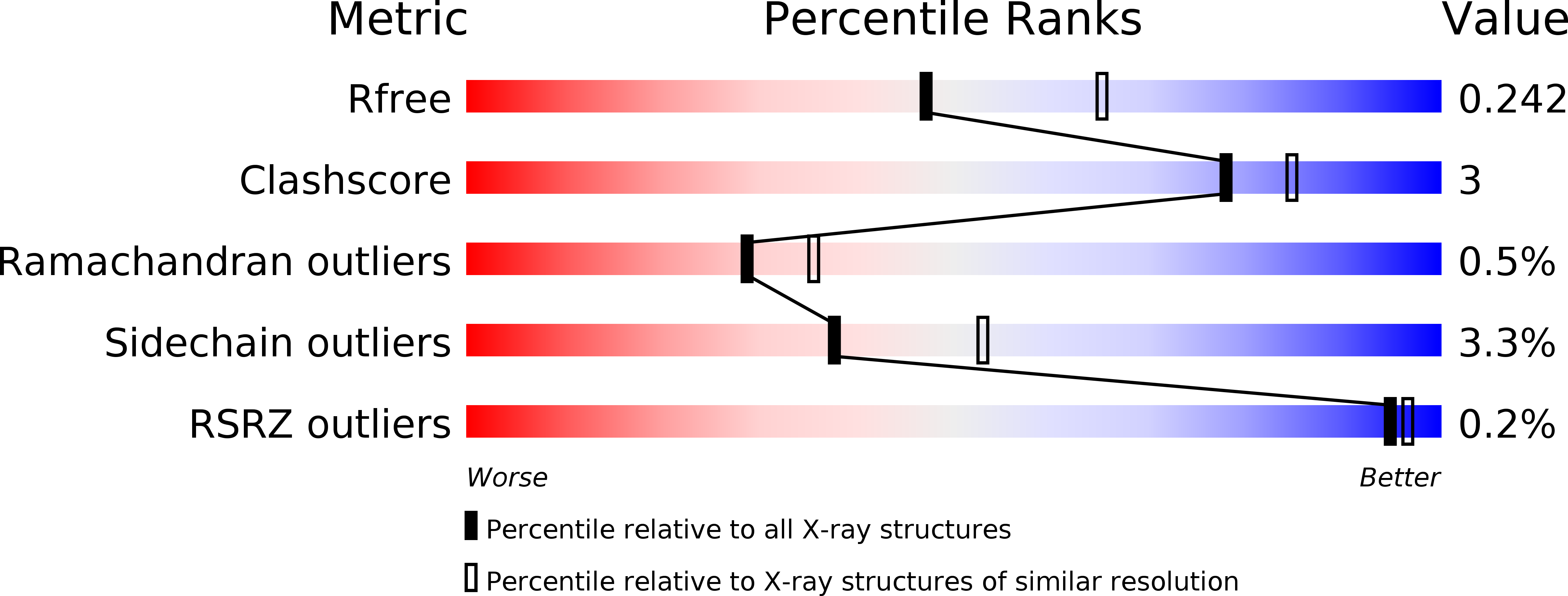
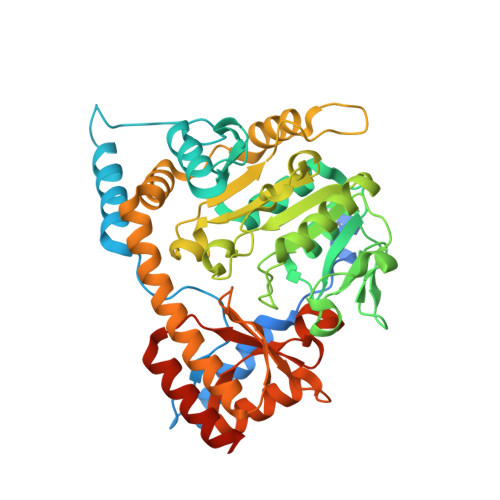
Structure of ALD1, a plant-specific homologue of the universal diaminopimelate aminotransferase enzyme of lysine biosynthesis.
Sobolev, V., Edelman, M., Dym, O., Unger, T., Albeck, S., Kirma, M., Galili, G.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 84-89
- PubMed: 23385743
- DOI: https://doi.org/10.1107/S1744309112050270
- Primary Citation of Related Structures:
4FL0 - PubMed Abstract:
Diaminopimelate aminotransferase (DAP-AT) is an enzyme in the lysine-biosynthesis pathway. Conversely, ALD1, a close homologue of DAP-AT in plants, uses lysine as a substrate in vitro. Both proteins require pyridoxal-5'-phosphate (PLP) for their activity. The structure of ALD1 from the flowering plant Arabidopsis thaliana (AtALD1) was solved at a resolution of 2.3 Å. Comparison of AtALD1 with the previously solved structure of A. thaliana DAP-AT (AtDAP-AT) revealed similar interactions with PLP despite sequence differences within the PLP-binding site. However, sequence differences between the binding site of AtDAP-AT for malate, a purported mimic of substrate binding, and the corresponding site in AtALD1 led to different interactions. This suggests that either the substrate itself, or the substrate-binding mode, differs in the two proteins, supporting the known in vitro findings.
Organizational Affiliation:
Department of Plant Sciences, Weizmann Institute of Science, Rehovot 76100, Israel. vladimir.sobolev@weizmann.ac.il