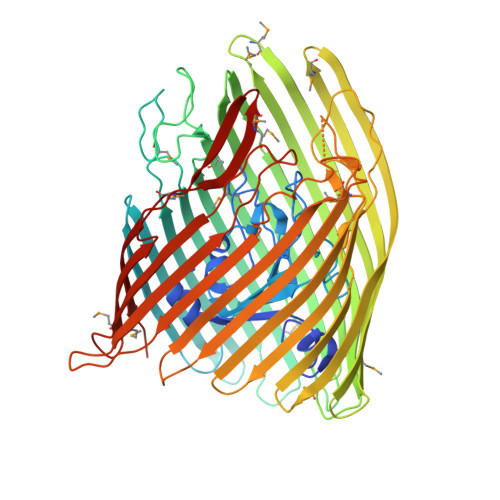
Structural engineering of a phage lysin that targets Gram-negative pathogens.
Lukacik, P., Barnard, T.J., Keller, P.W., Chaturvedi, K.S., Seddiki, N., Fairman, J.W., Noinaj, N., Kirby, T.L., Henderson, J.P., Steven, A.C., Hinnebusch, B.J., Buchanan, S.K.(2012) Proc Natl Acad Sci U S A 109: 9857-9862
- PubMed: 22679291
- DOI: https://doi.org/10.1073/pnas.1203472109
- Primary Citation of Related Structures:
4EPA, 4EPF, 4EPI, 4EXM - PubMed Abstract:
Bacterial pathogens are becoming increasingly resistant to antibiotics. As an alternative therapeutic strategy, phage therapy reagents containing purified viral lysins have been developed against gram-positive organisms but not against gram-negative organisms due to the inability of these types of drugs to cross the bacterial outer membrane. We solved the crystal structures of a Yersinia pestis outer membrane transporter called FyuA and a bacterial toxin called pesticin that targets this transporter. FyuA is a β-barrel membrane protein belonging to the family of TonB dependent transporters, whereas pesticin is a soluble protein with two domains, one that binds to FyuA and another that is structurally similar to phage T4 lysozyme. The structure of pesticin allowed us to design a phage therapy reagent comprised of the FyuA binding domain of pesticin fused to the N-terminus of T4 lysozyme. This hybrid toxin kills specific Yersinia and pathogenic E. coli strains and, importantly, can evade the pesticin immunity protein (Pim) giving it a distinct advantage over pesticin. Furthermore, because FyuA is required for virulence and is more common in pathogenic bacteria, the hybrid toxin also has the advantage of targeting primarily disease-causing bacteria rather than indiscriminately eliminating natural gut flora.
Organizational Affiliation:
Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892, USA.