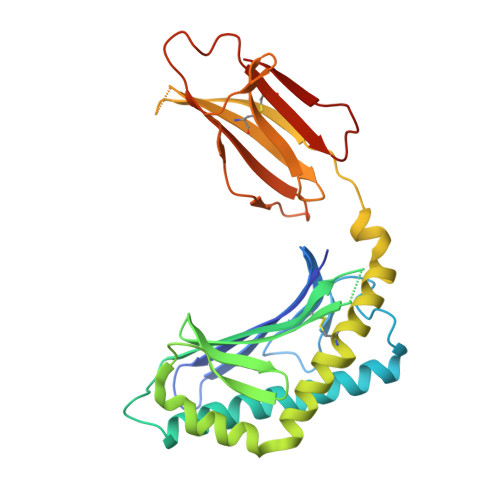
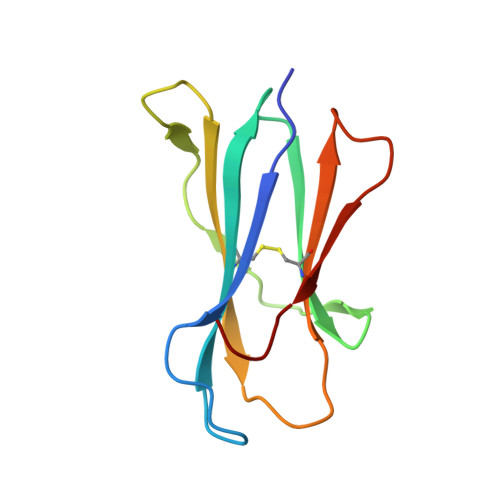
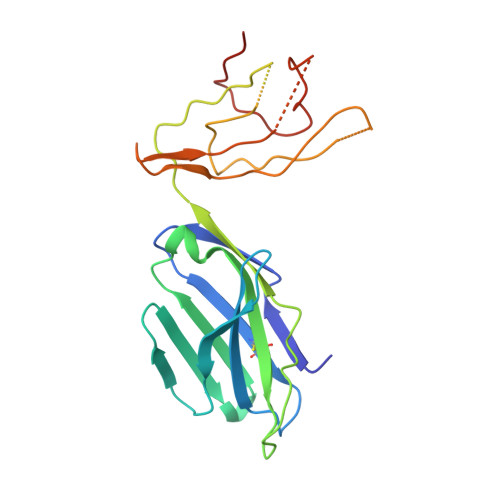
Type II natural killer T cells use features of both innate-like and conventional T cells to recognize sulfatide self antigens.
Girardi, E., Maricic, I., Wang, J., Mac, T.T., Iyer, P., Kumar, V., Zajonc, D.M.(2012) Nat Immunol 13: 851-856
- PubMed: 22820602
- DOI: https://doi.org/10.1038/ni.2371
- Primary Citation of Related Structures:
4ELK, 4ELM - PubMed Abstract:
Glycolipids presented by the major histocompatibility complex (MHC) class I homolog CD1d are recognized by natural killer T cells (NKT cells) characterized by either a semi-invariant T cell antigen receptor (TCR) repertoire (type I NKT cells or iNKT cells) or a relatively variable TCR repertoire (type II NKT cells). Here we describe the structure of a type II NKT cell TCR in complex with CD1d-lysosulfatide. Both TCR α-chains and TCR β-chains made contact with the CD1d molecule with a diagonal footprint, typical of MHC-TCR interactions, whereas the antigen was recognized exclusively with a single TCR chain, similar to the iNKT cell TCR. Type II NKT cell TCRs, therefore, recognize CD1d-sulfatide complexes by a distinct recognition mechanism characterized by the TCR-binding features of both iNKT cells and conventional peptide-reactive T cells.
Organizational Affiliation:
Division of Cell Biology, La Jolla Institute for Allergy & Immunology, La Jolla, California, USA.