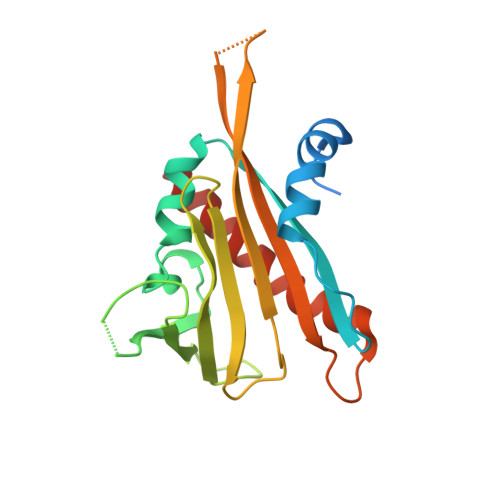
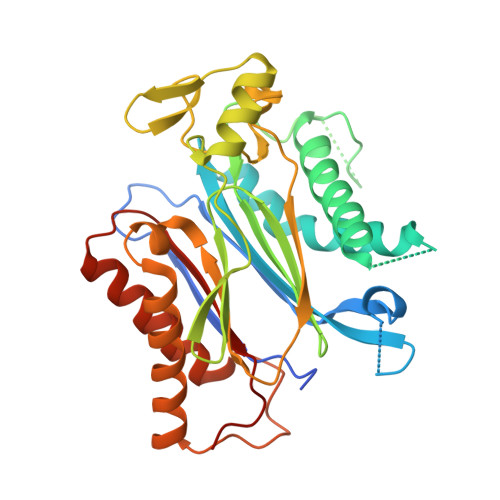
Complex Structures of the Abscisic Acid Receptor PYL3/RCAR13 Reveal a Unique Regulatory Mechanism
Zhang, X., Zhang, Q., Xin, Q., Yu, L., Wang, Z., Wu, W., Jiang, L., Wang, G., Tian, W., Deng, Z., Wang, Y., Liu, Z., Long, J., Gong, Z., Chen, Z.(2012) Structure 20: 780-790
- PubMed: 22579247
- DOI: https://doi.org/10.1016/j.str.2012.02.019
- Primary Citation of Related Structures:
3KL1, 3KLX, 3OJI, 4DS8, 4DSB, 4DSC - PubMed Abstract:
Abscisic acid (ABA) controls many physiological processes and mediates adaptive responses to abiotic stresses. The ABA signaling mechanisms for abscisic acid receptors PYR/PYL/RCAR (PYLs) were reported. However, it remains unclear whether the molecular mechanisms are suitable for other PYLs. Here, complex structures of PYL3 with (+)-ABA, pyrabactin and HAB1 are reported. An unexpected trans-homodimer intermediate observed in the crystal is confirmed in solution. ABA-bound PYL3 greatly promotes the generation of monomeric PYL3, which can excessively increase the efficiency of inhibiting PP2Cs. Structure-guided biochemical experiments show that Ser195 accounts for the key intermediate. Interestingly, pyrabactin binds to PYL3 in a distinct nonproductive mode with gate closure, which sheds light on the design of agonists and antagonists for abscisic acid receptors. According to different conformations of ligand-bound PYLs, the PYLs family can be divided into three subclasses, among which the trans-dimeric subclass, represented by PYL3, reveals a distinct regulatory mechanism.
Organizational Affiliation:
State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing 100193, China.