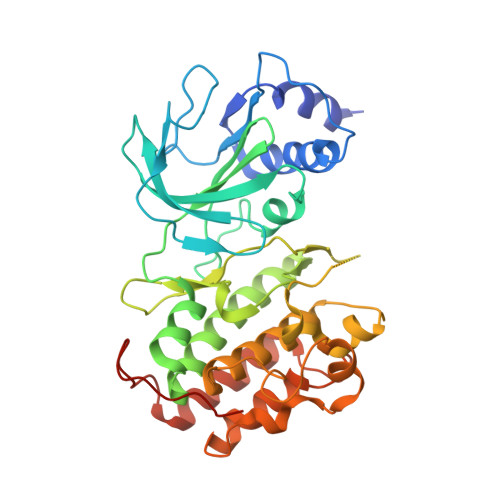
Structure of Nuclear Factor Kappa B-inducing kinase domain reveals a constitutively active conformation
Liu, J., Sudom, A., Min, X., Cao, Z., Gao, X., Ayres, M., Lee, F., Cao, P., Johnstone, S., Plotnikova, O., Walker, N., Chen, G., Wang, Z.(2012) J Biol Chem 287: 27326-27334
- PubMed: 22718757
- DOI: https://doi.org/10.1074/jbc.M112.366658
- Primary Citation of Related Structures:
4DN5 - PubMed Abstract:
NF-κB-inducing kinase (NIK) is a central component in the non-canonical NF-κB signaling pathway. Excessive NIK activity is implicated in various disorders, such as autoimmune conditions and cancers. Here, we report the first crystal structure of truncated human NIK in complex with adenosine 5'-O-(thiotriphosphate) at a resolution of 2.5 Å. This truncated protein is a catalytically active construct, including an N-terminal extension of 60 residues prior to the kinase domain, the kinase domain, and 20 residues afterward. The structure reveals that the NIK kinase domain assumes an active conformation in the absence of any phosphorylation. Analysis of the structure uncovers a unique role for the N-terminal extension sequence, which stabilizes helix αC in the active orientation and keeps the kinase domain in the catalytically competent conformation. Our findings shed light on the long-standing debate over whether NIK is a constitutively active kinase. They also provide a molecular basis for the recent observation of gain-of-function activity for an N-terminal deletion mutant (ΔN324) of NIK, leading to constitutive non-canonical NF-κB signaling with enhanced B-cell adhesion and apoptosis resistance.
Organizational Affiliation:
Amgen Inc., South San Francisco, California 94080, USA.