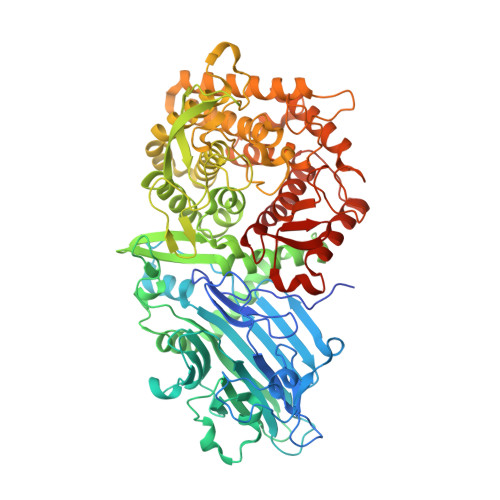
A Novel Beta-Xylosidase Structure from Geobacillus Thermoglucosidasius: The First Crystal Structure of a Glycoside Hydrolase Family Gh52 Enzyme Reveals Unpredicted Similarity to Other Glycoside Hydrolase Folds
Espina, G., Eley, K., Pompidor, G., Schneider, T.R., Crennell, S.J., Danson, M.J.(2014) Acta Crystallogr D Biol Crystallogr 70: 1366
- PubMed: 24816105
- DOI: https://doi.org/10.1107/S1399004714002788
- Primary Citation of Related Structures:
4C1O, 4C1P - PubMed Abstract:
Geobacillus thermoglucosidasius is a thermophilic bacterium that is able to ferment both C6 and C5 sugars to produce ethanol. During growth on hemicellulose biomass, an intracellular β-xylosidase catalyses the hydrolysis of xylo-oligosaccharides to the monosaccharide xylose, which can then enter the pathways of central metabolism. The gene encoding a G. thermoglucosidasius β-xylosidase belonging to CAZy glycoside hydrolase family GH52 has been cloned and expressed in Escherichia coli. The recombinant enzyme has been characterized and a high-resolution (1.7 Å) crystal structure has been determined, resulting in the first reported structure of a GH52 family member. A lower resolution (2.6 Å) structure of the enzyme-substrate complex shows the positioning of the xylobiose substrate to be consistent with the proposed retaining mechanism of the family; additionally, the deep cleft of the active-site pocket, plus the proximity of the neighbouring subunit, afford an explanation for the lack of catalytic activity towards the polymer xylan. Whilst the fold of the G. thermoglucosidasius β-xylosidase is completely different from xylosidases in other CAZy families, the enzyme surprisingly shares structural similarities with other glycoside hydrolases, despite having no more than 13% sequence identity.
Organizational Affiliation:
Centre for Extremophile Research, Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, England.