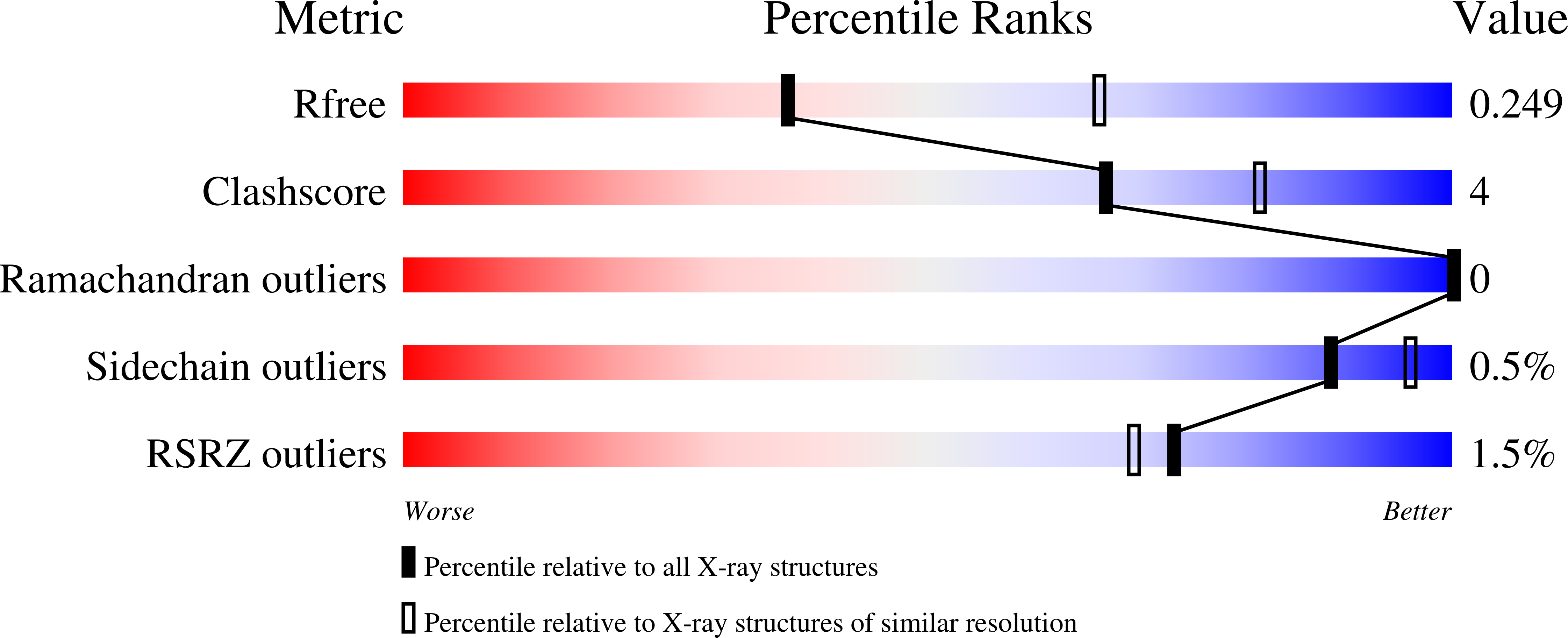
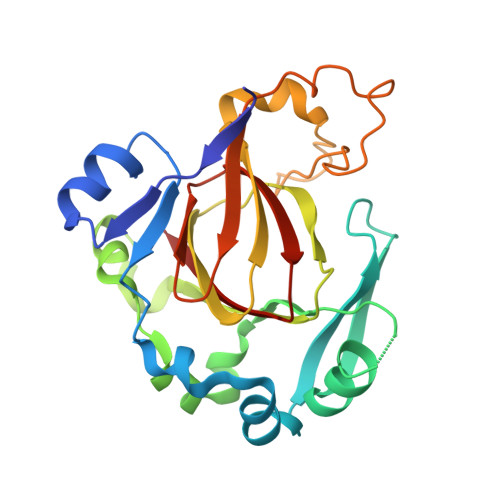
Crystal Structure of Jmjd5 Domain of Human Lysine- Specific Demethylase 8 (Kdm8) in Complex with N- Oxalylglycine (Nog)
Vollmar, M., Johansson, C., Krojer, T., Canning, P., Allerston, C., Gadhave, A., von Delft, F., Bountra, C., Arrowsmith, C.H., Weigelt, J., Edwards, A., Oppermann, U.To be published.