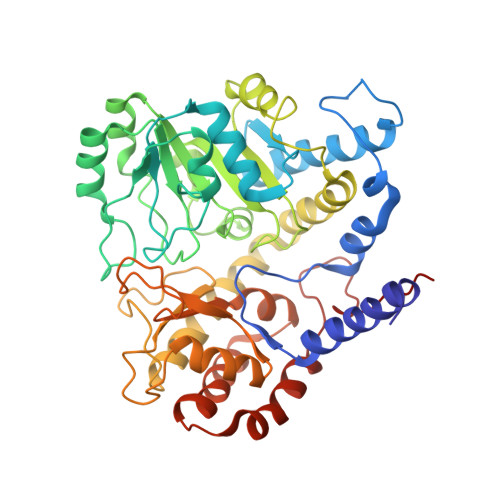
The Crystal Structure of Archaeal Serine Hydroxymethyltransferase Reveals Idiosyncratic Features Likely Required to Withstand High Temperatures.
Angelucci, F., Morea, V., Angelaccio, S., Saccoccia, F., Contestabile, R., Ilari, A.(2014) Proteins 82: 3437
- PubMed: 25257552
- DOI: https://doi.org/10.1002/prot.24697
- Primary Citation of Related Structures:
4BHD, 4UQV - PubMed Abstract:
Serine hydroxymethyltransferases (SHMTs) play an essential role in one-carbon unit metabolism and are used in biomimetic reactions. We determined the crystal structure of free (apo) and pyridoxal-5'-phosphate-bound (holo) SHMT from Methanocaldococcus jannaschii, the first from a hyperthermophile, from the archaea domain of life and that uses H₄MPT as a cofactor, at 2.83 and 3.0 Å resolution, respectively. Idiosyncratic features were observed that are likely to contribute to structure stabilization. At the dimer interface, the C-terminal region folds in a unique fashion with respect to SHMTs from eubacteria and eukarya. At the active site, the conserved tyrosine does not make a cation-π interaction with an arginine like that observed in all other SHMT structures, but establishes an amide-aromatic interaction with Asn257, at a different sequence position. This asparagine residue is conserved and occurs almost exclusively in (hyper)thermophile SHMTs. This led us to formulate the hypothesis that removal of frustrated interactions (such as the Arg-Tyr cation-π interaction occurring in mesophile SHMTs) is an additional strategy of adaptation to high temperature. Both peculiar features may be tested by designing enzyme variants potentially endowed with improved stability for applications in biomimetic processes.
Organizational Affiliation:
Department of Life, Health and Environmental Sciences, University of L'Aquila, P.le Salvatore Tommasi 1, L'Aquila, Italy.