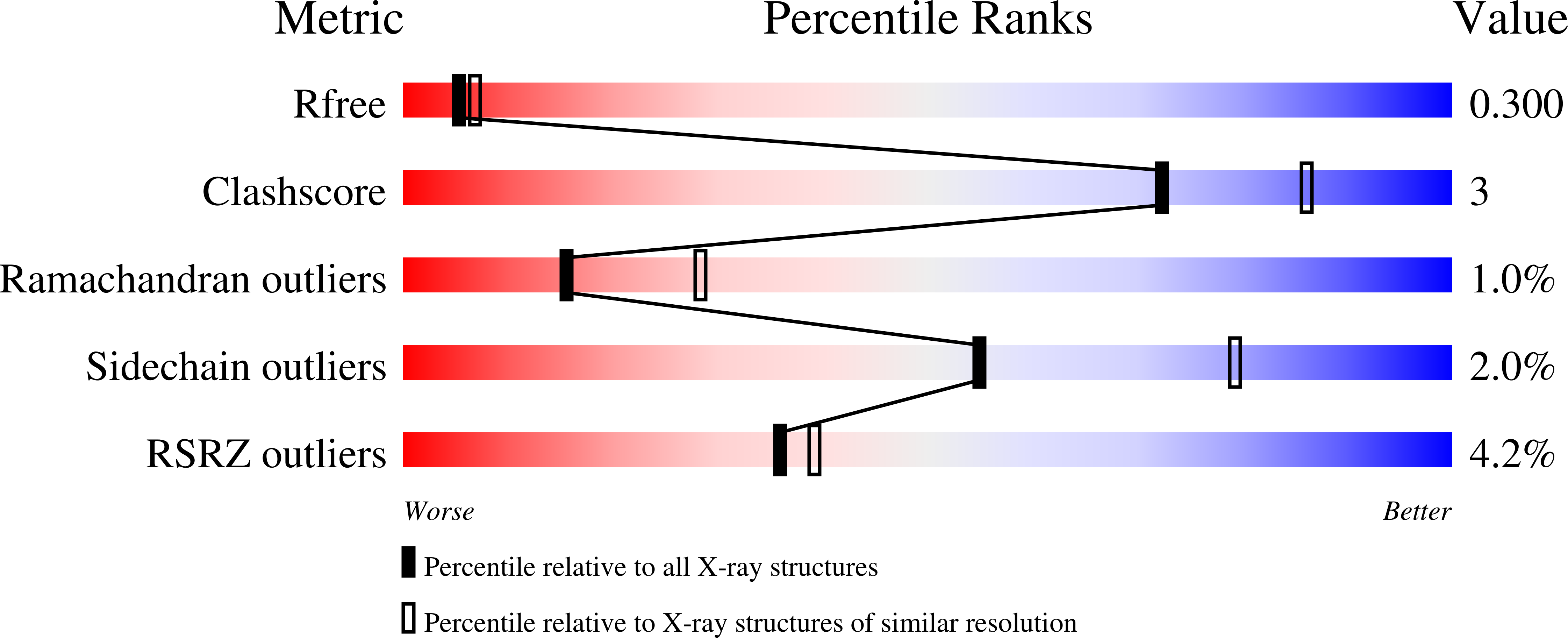
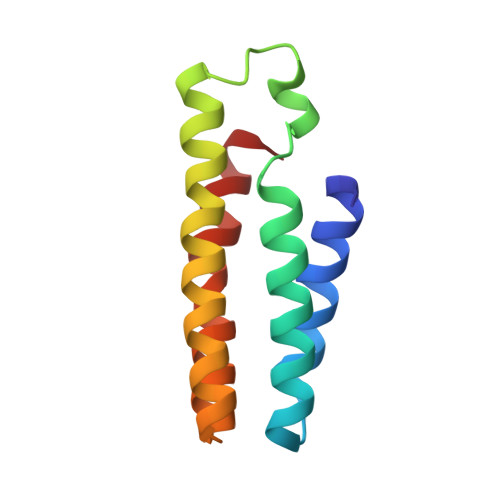
A designed supramolecular protein assembly with in vivo enzymatic activity.
Song, W.J., Tezcan, F.A.(2014) Science 346: 1525-1528
- PubMed: 25525249
- DOI: https://doi.org/10.1126/science.1259680
- Primary Citation of Related Structures:
4U9D, 4U9E - PubMed Abstract:
The generation of new enzymatic activities has mainly relied on repurposing the interiors of preexisting protein folds because of the challenge in designing functional, three-dimensional protein structures from first principles. Here we report an artificial metallo-β-lactamase, constructed via the self-assembly of a structurally and functionally unrelated, monomeric redox protein into a tetrameric assembly that possesses catalytic zinc sites in its interfaces. The designed metallo-β-lactamase is functional in the Escherichia coli periplasm and enables the bacteria to survive treatment with ampicillin. In vivo screening of libraries has yielded a variant that displays a catalytic proficiency [(k(cat)/K(m))/k(uncat)] for ampicillin hydrolysis of 2.3 × 10(6) and features the emergence of a highly mobile loop near the active site, a key component of natural β-lactamases to enable substrate interactions.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of California, San Diego, La Jolla, CA 92093-0356, USA.