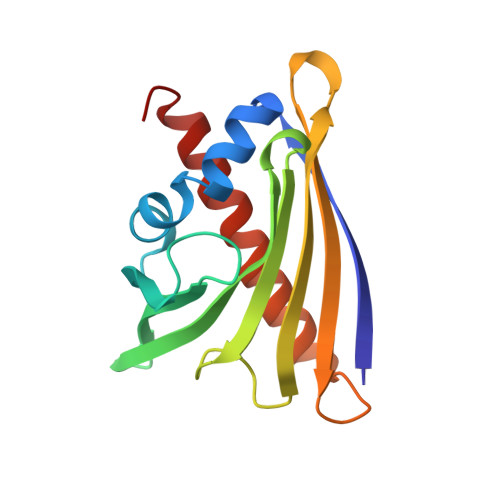
Crystallographic and CD probing of ligand-induced conformational changes in a plant PR-10 protein.
Sliwiak, J., Dolot, R., Michalska, K., Szpotkowski, K., Bujacz, G., Sikorski, M., Jaskolski, M.(2016) J Struct Biol 193: 55-66
- PubMed: 26644353
- DOI: https://doi.org/10.1016/j.jsb.2015.11.008
- Primary Citation of Related Structures:
4RYV, 4Y31, 5C9Y - PubMed Abstract:
Plant pathogenesis-related class 10 (PR-10) proteins are a family of abundant proteins initially identified as elements of the plant defense system. The key structural feature suggesting PR-10 functionality is a huge hydrophobic cavity created in the protein interior by a scaffold composed of an extended β-sheet wrapped around a long and flexible C-terminal α-helix. Several crystallographic and NMR studies have shown that the cavity can accommodate a variety of small molecule ligands, including phytohormones. The article describes ∼1.3 Å resolution crystal structures of a Lupinus luteus PR-10 isoform LlPR-10.1A, in its free form and in complex with trans-zeatin, a naturally occurring plant hormone belonging to the cytokinin group. Moreover we present the structure of the same protein where the saturation with zeatin is not complete. This set of three crystal structures allows us to track the structural adaptation of the protein upon trans-zeatin docking, as well as the sequence of the ligand-binding events, step-by-step. In addition, titration of LlPR-10.1A with trans-zeatin monitored in solution by CD spectra, confirmed the pattern of structural adaptations deduced from the crystallographic studies. The ligand-biding mode shows no similarity to other zeatin complexes of PR-10 proteins. The present work, which describes the first atomic models of the same PR-10 protein with and without a physiological ligand, reveals that the conformation of LlPR-10.1A undergoes a significant structural rearrangement upon trans-zeatin binding.
Organizational Affiliation:
Center for Biocrystallographic Research, Institute of Bioorganic Chemistry, Polish Academy of Sciences, Poznan, Poland.