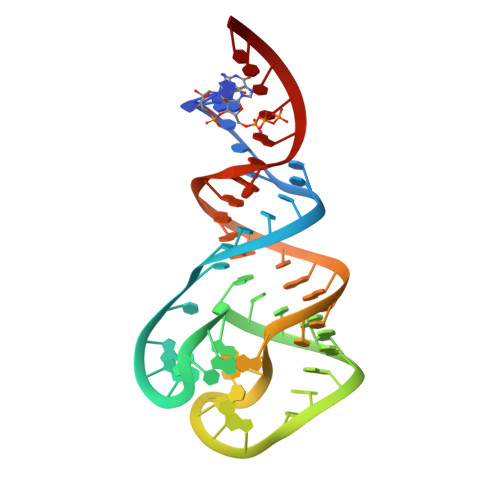
T box RNA decodes both the information content and geometry of tRNA to affect gene expression.
Grigg, J.C., Chen, Y., Grundy, F.J., Henkin, T.M., Pollack, L., Ke, A.(2013) Proc Natl Acad Sci U S A 110: 7240-7245
- PubMed: 23589841
- DOI: https://doi.org/10.1073/pnas.1222214110
- Primary Citation of Related Structures:
4JRC - PubMed Abstract:
The T box leader sequence is an RNA element that controls gene expression by binding directly to a specific tRNA and sensing its aminoacylation state. This interaction controls expression of amino acid-related genes in a negative feedback loop. The T box RNA structure is highly conserved, but its tRNA binding mechanism is only partially understood. Known sequence elements are the specifier sequence, which recognizes the tRNA anticodon, and the antiterminator bulge, which base pairs with the tRNA acceptor end. Here, we reveal the crucial function of the highly conserved stem I distal region in tRNA recognition and report its 2.65-Å crystal structure. The apex of this region contains an intricately woven loop-loop interaction between two conserved motifs, the Adenine-guanine (AG) bulge and the distal loop. This loop-loop structure presents a base triple on its surface that is optimally positioned for base-stacking interactions. Mutagenesis, cross-linking, and small-angle X-ray scattering data demonstrate that the apical base triple serves as a binding platform to dock the tRNA D- and T-loops. Strikingly, the binding platform strongly resembles the D- and T-loop binding elements from RNase P and the ribosome exit site, suggesting that this loop-loop structure may represent a widespread tRNA recognition platform. We propose a two-checkpoint molecular ruler model for tRNA decoding in which the information content of tRNA is first examined through specifier sequence-anticodon interaction, and the length of the tRNA anticodon arm is then measured by the distal loop-loop platform. When both conditions are met, tRNA is secured, and its aminoacylation state is sensed.
Organizational Affiliation:
Department of Molecular Biology and Genetics, Cornell University, Ithaca, NY 14850, USA.