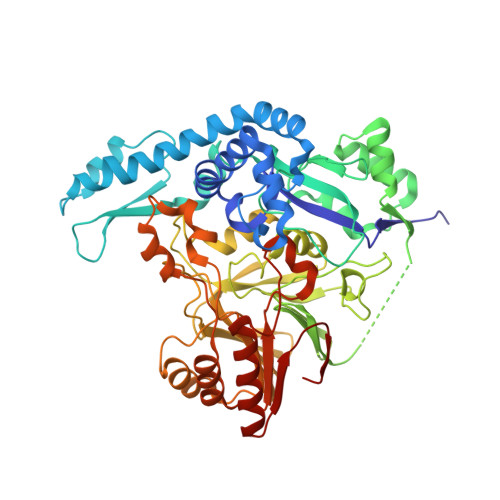
Structure of a Dinuclear Iron Cluster-Containing beta-Hydroxylase Active in Antibiotic Biosynthesis.
Makris, T.M., Knoot, C.J., Wilmot, C.M., Lipscomb, J.D.(2013) Biochemistry 52: 6662-6671
- PubMed: 23980641
- DOI: https://doi.org/10.1021/bi400845b
- Primary Citation of Related Structures:
4JO0 - PubMed Abstract:
A family of dinuclear iron cluster-containing oxygenases that catalyze β-hydroxylation tailoring reactions in natural product biosynthesis by nonribosomal peptide synthetase (NRPS) systems was recently described [Makris, T. M., Chakrabarti, M., Münck, E., and Lipscomb, J. D. (2010) Proc. Natl. Acad. Sci. U.S.A. 107, 15391-15396]. Here, the 2.17 Å X-ray crystal structure of the archetypal enzyme from the family, CmlA, is reported. CmlA catalyzes β-hydroxylation of l-p-aminophenylalanine during chloramphenicol biosynthesis. The fold of the N-terminal domain of CmlA is unlike any previously reported, but the C-terminal domain has the αββα fold of the metallo-β-lactamase (MBL) superfamily. The diiron cluster bound in the C-terminal domain is coordinated by an acetate, three His residues, two Asp residues, one Glu residue, and a bridging oxo moiety. One of the Asp ligands forms an unusual monodentate bridge. No other oxygen-activating diiron enzyme utilizes this ligation or the MBL protein fold. The N-terminal domain facilitates dimerization, but using computational docking and a sequence-based structural comparison to homologues, we hypothesize that it likely serves additional roles in NRPS recognition and the regulation of O2 activation.
Organizational Affiliation:
Department of Biochemistry, Molecular Biology and Biophysics and Center for Metals in Biocatalysis, University of Minnesota , Minneapolis, Minnesota 55455, United States.