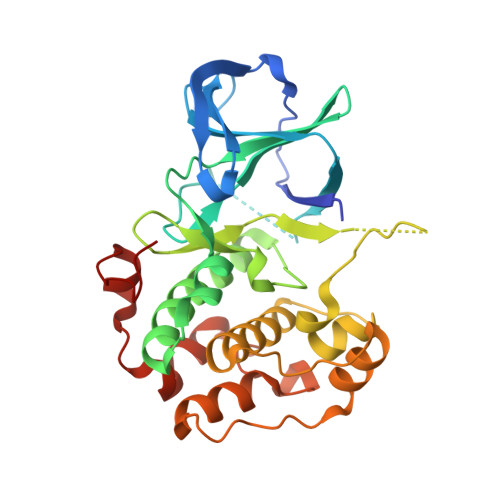
Identification of quercitrin as an inhibitor of the p90 S6 ribosomal kinase (RSK): structure of its complex with the N-terminal domain of RSK2 at 1.8 A resolution.
Derewenda, U., Artamonov, M., Szukalska, G., Utepbergenov, D., Olekhnovich, N., Parikh, H.I., Kellogg, G.E., Somlyo, A.V., Derewenda, Z.S.(2013) Acta Crystallogr D Biol Crystallogr 69: 266-275
- PubMed: 23385462
- DOI: https://doi.org/10.1107/S0907444912045520
- Primary Citation of Related Structures:
4GUE - PubMed Abstract:
Members of the RSK family of kinases constitute attractive targets for drug design, but a lack of structural information regarding the mechanism of selective inhibitors impedes progress in this field. The crystal structure of the N-terminal kinase domain (residues 45-346) of mouse RSK2, or RSK2(NTKD), has recently been described in complex with one of only two known selective inhibitors, a rare naturally occurring flavonol glycoside, kaempferol 3-O-(3'',4''-di-O-acetyl-α-L-rhamnopyranoside), known as SL0101. Based on this structure, it was hypothesized that quercitrin (quercetin 3-O-α-L-rhamnopyranoside), a related but ubiquitous and inexpensive compound, might also act as an RSK inhibitor. Here, it is demonstrated that quercitrin binds to RSK2(NTKD) with a dissociation constant (K(d)) of 5.8 µM as determined by isothermal titration calorimetry, and a crystal structure of the binary complex at 1.8 Å resolution is reported. The crystal structure reveals a very similar mode of binding to that recently reported for SL0101. Closer inspection shows a number of small but significant differences that explain the slightly higher K(d) for quercitrin compared with SL0101. It is also shown that quercitrin can effectively substitute for SL0101 in a biological assay, in which it significantly suppresses the contractile force in rabbit pulmonary artery smooth muscle in response to Ca(2+).
Organizational Affiliation:
Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville, VA 22908-0736, USA.