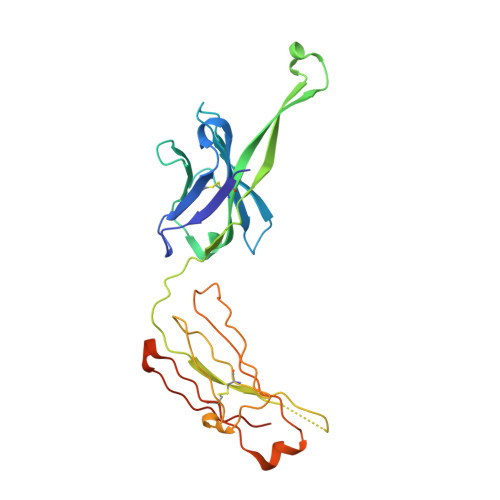
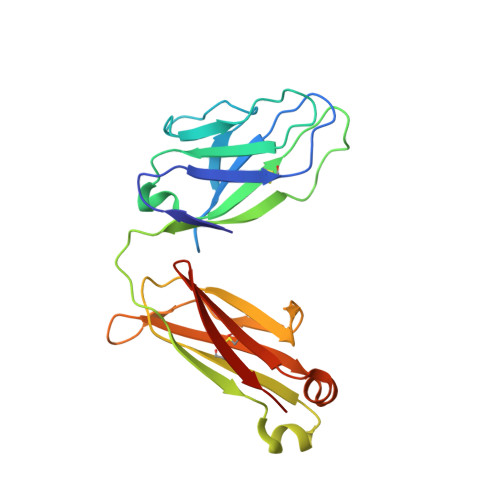
Complex-type N-glycan recognition by potent broadly neutralizing HIV antibodies.
Mouquet, H., Scharf, L., Euler, Z., Liu, Y., Eden, C., Scheid, J.F., Halper-Stromberg, A., Gnanapragasam, P.N., Spencer, D.I., Seaman, M.S., Schuitemaker, H., Feizi, T., Nussenzweig, M.C., Bjorkman, P.J.(2012) Proc Natl Acad Sci U S A 109: E3268-E3277
- PubMed: 23115339
- DOI: https://doi.org/10.1073/pnas.1217207109
- Primary Citation of Related Structures:
4FQ1, 4FQ2, 4FQC, 4FQQ - PubMed Abstract:
Broadly neutralizing HIV antibodies (bNAbs) can recognize carbohydrate-dependent epitopes on gp120. In contrast to previously characterized glycan-dependent bNAbs that recognize high-mannose N-glycans, PGT121 binds complex-type N-glycans in glycan microarrays. We isolated the B-cell clone encoding PGT121, which segregates into PGT121-like and 10-1074-like groups distinguished by sequence, binding affinity, carbohydrate recognition, and neutralizing activity. Group 10-1074 exhibits remarkable potency and breadth but no detectable binding to protein-free glycans. Crystal structures of unliganded PGT121, 10-1074, and their likely germ-line precursor reveal that differential carbohydrate recognition maps to a cleft between complementarity determining region (CDR)H2 and CDRH3. This cleft was occupied by a complex-type N-glycan in a "liganded" PGT121 structure. Swapping glycan contact residues between PGT121 and 10-1074 confirmed their importance for neutralization. Although PGT121 binds complex-type N-glycans, PGT121 recognized high-mannose-only HIV envelopes in isolation and on virions. As HIV envelopes exhibit varying proportions of high-mannose- and complex-type N-glycans, these results suggest promiscuous carbohydrate interactions, an advantageous adaptation ensuring neutralization of all viruses within a given strain.
Organizational Affiliation:
Laboratory of Molecular Immunology, The Rockefeller University, New York, NY 10021, USA. hmouquet@rockefeller.edu