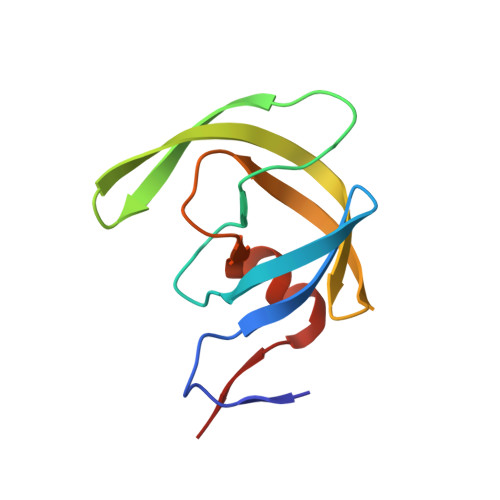
Higher Desolvation Energy Reduces Molecular Recognition in Multi-Drug Resistant HIV-1 Protease.
Wang, Y., Dewdney, T.G., Liu, Z., Reiter, S.J., Brunzelle, J.S., Kovari, I.A., Kovari, L.C.(2012) Biology (Basel) 1: 81-93
- PubMed: 24832048
- DOI: https://doi.org/10.3390/biology1010081
- Primary Citation of Related Structures:
4FAE, 4FAF - PubMed Abstract:
Designing HIV-1 protease inhibitors that overcome drug-resistance is still a challenging task. In this study, four clinical isolates of multi-drug resistant HIV-1 proteases that exhibit resistance to all the US FDA-approved HIV-1 protease inhibitors and also reduce the substrate recognition ability were examined. A multi-drug resistant HIV-1 protease isolate, MDR 769, was co-crystallized with the p2/NC substrate and the mutated CA/p2 substrate, CA/p2 P1'F. Both substrates display different levels of molecular recognition by the wild-type and multi-drug resistant HIV-1 protease. From the crystal structures, only limited differences can be identified between the wild-type and multi-drug resistant protease. Therefore, a wild-type HIV-1 protease and four multi-drug resistant HIV-1 proteases in complex with the two peptides were modeled based on the crystal structures and examined during a 10 ns-molecular dynamics simulation. The simulation results reveal that the multi-drug resistant HIV-1 proteases require higher desolvation energy to form complexes with the peptides. This result suggests that the desolvation of the HIV-1 protease active site is an important step of protease-ligand complex formation as well as drug resistance. Therefore, desolvation energy could be considered as a parameter in the evaluation of future HIV-1 protease inhibitor candidates.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Wayne State University School of Medicine, Detroit, MI 48201, USA. yowang@med.wayne.edu.