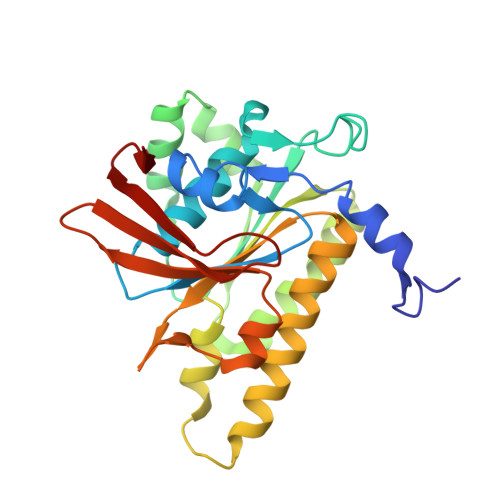
Structure of a hexameric form of RadA recombinase from Methanococcus voltae.
Du, L., Luo, Y.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 511-516
- PubMed: 22691778
- DOI: https://doi.org/10.1107/S1744309112010226
- Primary Citation of Related Structures:
4DC9 - PubMed Abstract:
Archaeal RadA proteins are close homologues of eukaryal Rad51 and DMC1 proteins and are remote homologues of bacterial RecA proteins. For the repair of double-stranded breaks in DNA, these recombinases promote a pivotal strand-exchange reaction between homologous single-stranded and double-stranded DNA substrates. This DNA-repair function also plays a key role in the resistance of cancer cells to chemotherapy and radiotherapy and in the resistance of bacterial cells to antibiotics. A hexameric form of a truncated Methanococcus voltae RadA protein devoid of its small N-terminal domain has been crystallized. The RadA hexamers further assemble into two-ringed assemblies. Similar assemblies can be observed in the crystals of Pyrococcus furiosus RadA and Homo sapiens DMC1. In all of these two-ringed assemblies the DNA-interacting L1 region of each protomer points inward towards the centre, creating a highly positively charged locus. The electrostatic characteristics of the central channels can be utilized in the design of novel recombinase inhibitors.
Organizational Affiliation:
Department of Biochemistry, University of Saskatchewan, 107 Wiggins Road, Suite A3, Saskatoon, Sasktchewan S7N 5E5, Canada.