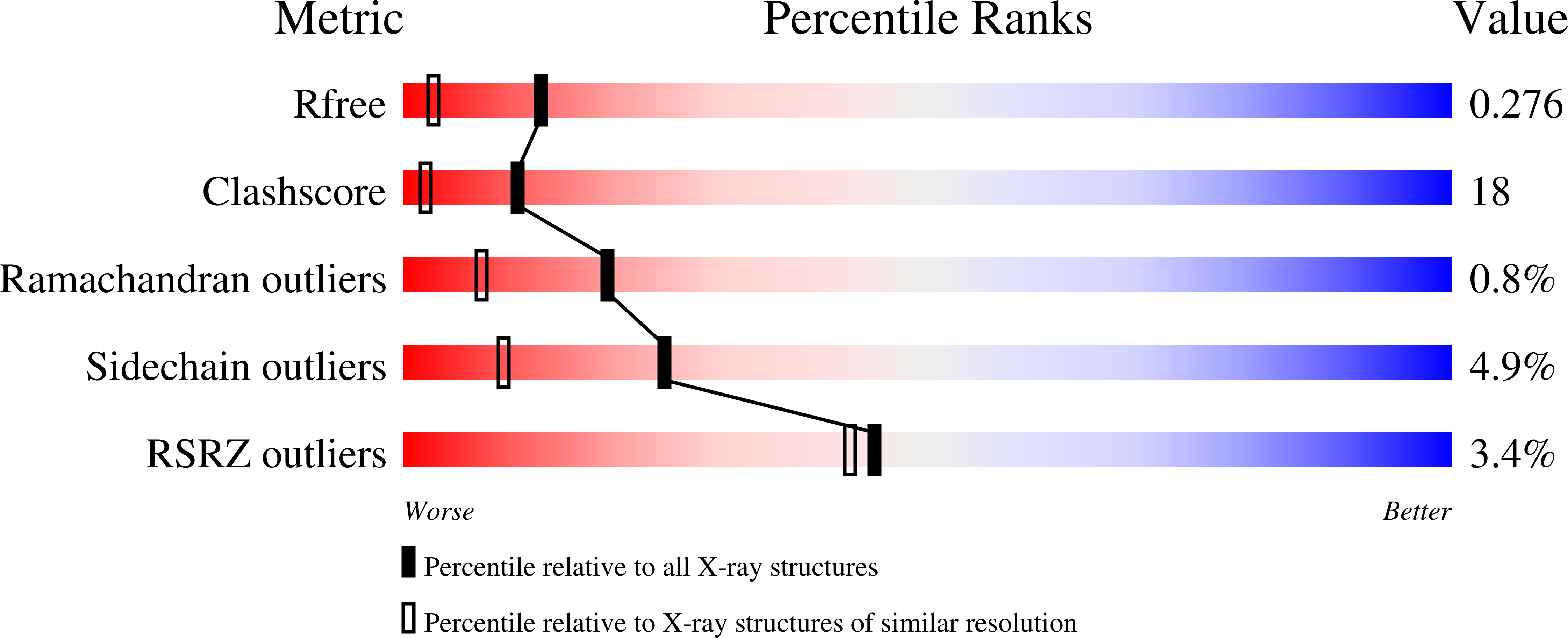
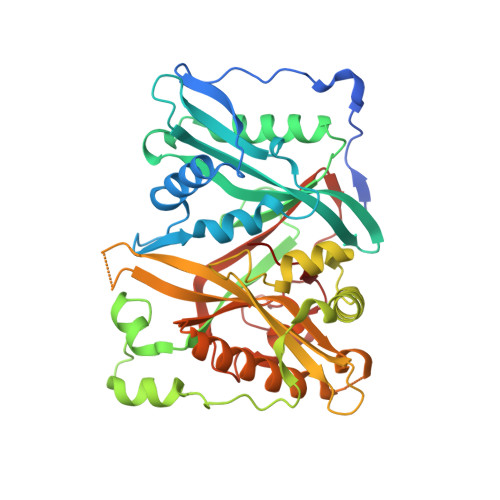
N-Myristoyltransferase is a Cell Wall Target in Aspergillus Fumigatus.
Fang, W., Robinson, D.A., Raimi, O.G., Blair, D.E., Harrison, J.R., Lockhart, D.E.A., Torrie, L.S., Ruda, G.F., Wyatt, P.G., Gilbert, I.H., Van Aalten, D.M.F.(2015) ACS Chem Biol 10: 1425
- PubMed: 25706802
- DOI: https://doi.org/10.1021/cb5008647
- Primary Citation of Related Structures:
4CAV, 4CAW, 4CAX - PubMed Abstract:
Treatment of filamentous fungal infections relies on a limited repertoire of antifungal agents. Compounds possessing novel modes of action are urgently required. N-myristoylation is a ubiquitous modification of eukaryotic proteins. The enzyme N-myristoyltransferase (NMT) has been considered a potential therapeutic target in protozoa and yeasts. Here, we show that the filamentous fungal pathogen Aspergillus fumigatus possesses an active NMT enzyme that is essential for survival. Surprisingly, partial repression of the gene revealed downstream effects of N-myristoylation on cell wall morphology. Screening a library of inhibitors led to the discovery of a pyrazole sulphonamide compound that inhibits the enzyme and is fungicidal under partially repressive nmt conditions. Together with a crystallographic complex showing the inhibitor binding in the peptide substrate pocket, we provide evidence of NMT being a potential drug target in A. fumigatus.
Organizational Affiliation:
†Division of Molecular Microbiology, ‡Division of Biological Chemistry and Drug Discovery, §MRC Protein Phosphorylation and Ubiquitylation Unit, College of Life Sciences, University of Dundee, Dundee, DD1 5EH, United Kingdom.