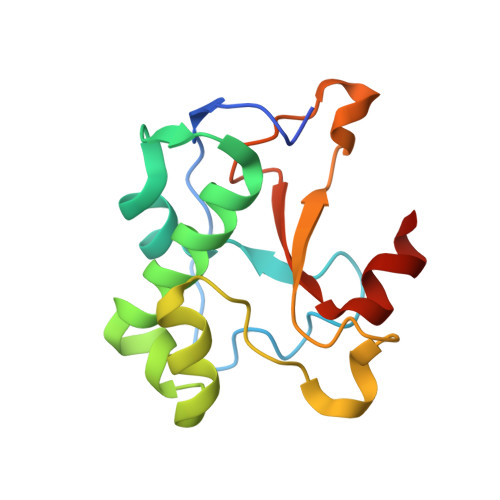
A New Insight Into the Zinc-Dependent DNA-Cleavage by the Colicin E7 Nuclease: A Crystallographic and Computational Study.
Czene, A., Toth, E., Nemeth, E., Otten, H., Poulsen, J.N., Christensen, H.E.M., Rulisek, L., Nagata, K., Larsen, S., Gyurcsik, B.(2014) Metallomics 6: 2090
- PubMed: 25179124
- DOI: https://doi.org/10.1039/c4mt00195h
- Primary Citation of Related Structures:
3ZFK - PubMed Abstract:
The nuclease domain of colicin E7 metallonuclease (NColE7) contains its active centre at the C-terminus. The mutant ΔN4-NColE7-C* - where the four N-terminal residues including the positively charged K446, R447 and K449 are replaced with eight residues from the GST tag - is catalytically inactive. The crystal structure of this mutant demonstrates that its overall fold is very similar to that of the native NColE7 structure. This implicates the stabilizing effect of the remaining N-terminal sequence on the structure of the C-terminal catalytic site and the essential role of the deleted residues in the mechanism of the catalyzed reaction. Complementary QM/MM calculations on the protein-DNA complexes support the less favourable cleavage by the mutant protein than by NColE7. Furthermore, a water molecule as a possible ligand for the Zn(2+)-ion is proposed to play a role in the catalytic process. These results suggest that the mechanism of the Zn(2+)-containing HNH nucleases needs to be further studied and discussed.
Organizational Affiliation:
MTA-SZTE Bioinorganic Chemistry Research Group, Dóm tér 7, H-6720 Szeged, Hungary.