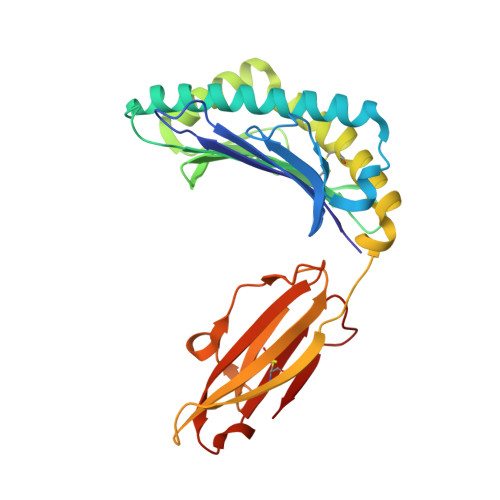
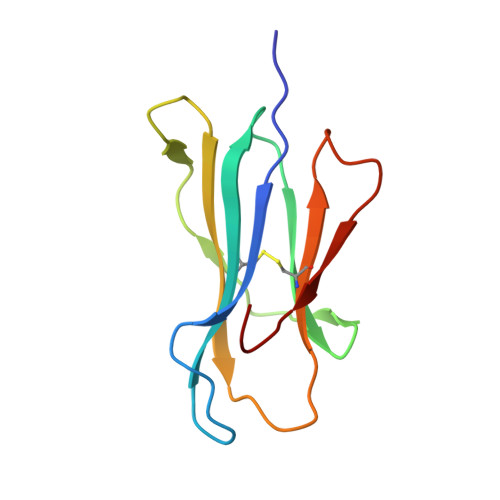
Switching and emergence of CTL epitopes in HIV-1 infection
Han, C., Kawana-tachikawa, A., Shimizu, A., Zhu, D., Nakamura, H., Adachi, E., Kikuchi, T., Koga, M., Koibuchi, T., Gao, G.F., Sato, Y., Yamagata, A., Martin, E., Fukai, S., Brumme, Z.L., Iwamoto, A.(2014) Retrovirology 11: 38-38
- PubMed: 24886641
- DOI: https://doi.org/10.1186/1742-4690-11-38
- Primary Citation of Related Structures:
3WL9, 3WLB - PubMed Abstract:
Human Leukocyte Antigen (HLA) class I restricted Cytotoxic T Lymphocytes (CTLs) exert substantial evolutionary pressure on HIV-1, as evidenced by the reproducible selection of HLA-restricted immune escape mutations in the viral genome. An escape mutation from tyrosine to phenylalanine at the 135th amino acid (Y135F) of the HIV-1 nef gene is frequently observed in patients with HLA-A*24:02, an HLA Class I allele expressed in ~70% of Japanese persons. The selection of CTL escape mutations could theoretically result in the de novo creation of novel epitopes, however, the extent to which such dynamic "CTL epitope switching" occurs in HIV-1 remains incompletely known.
Organizational Affiliation:
Division of Infectious Diseases, Advanced Clinical Research Center, the Institute of Medical Science, the University of Tokyo, 4-6-1 Shirokanedai, Minato-ku, Tokyo 108-8639, Japan. aikichi@ra3.so-net.ne.jp.