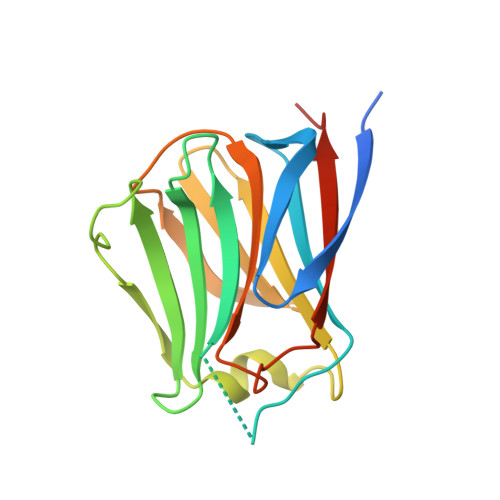
Conformational change of a unique sequence in a fungal galectin from Agrocybe cylindracea controls glycan ligand-binding specificity.
Kuwabara, N., Hu, D., Tateno, H., Makyio, H., Hirabayashi, J., Kato, R.(2013) FEBS Lett 587: 3620-3625
- PubMed: 24036446
- DOI: https://doi.org/10.1016/j.febslet.2013.08.046
- Primary Citation of Related Structures:
3WG1, 3WG2, 3WG3, 3WG4 - PubMed Abstract:
A fungal galectin from Agrocybe cylindracea (ACG) exhibits broad binding specificity for β-galactose-containing glycans. We determined the crystal structures of wild-type ACG and the N46A mutant, with and without glycan ligands. From these structures and a saccharide-binding analysis of the N46A mutant, we revealed that a conformational change of a unique insertion sequence containing Asn46 provides two binding modes for ACG, and thereby confers broad binding specificity. We propose that the unique sequence provides these two distinct glycan-binding modes by an induced-fit mechanism.
Organizational Affiliation:
Structural Biology Research Center, Photon Factory, Institute of Materials Structure Science, High Energy Accelerator Research Organization (KEK), 1-1 Oho, Tsukuba, Ibaraki 305-0801, Japan.