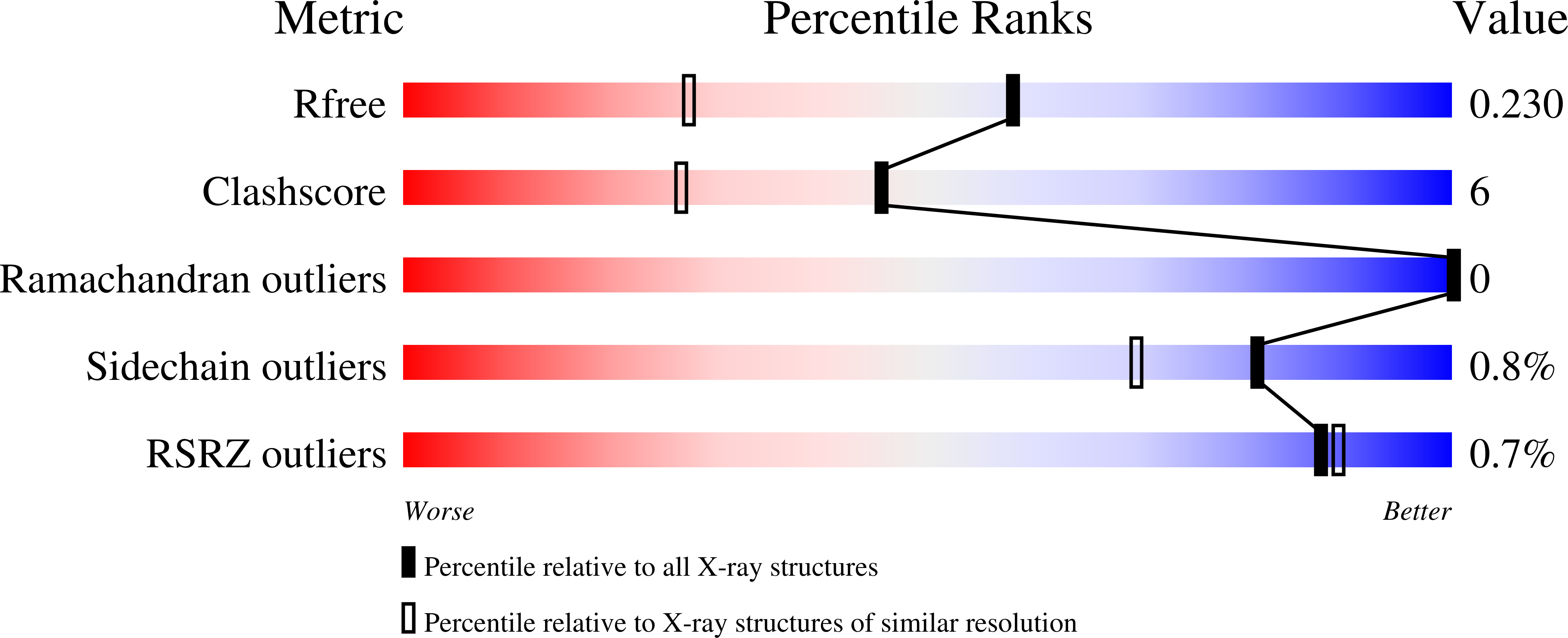
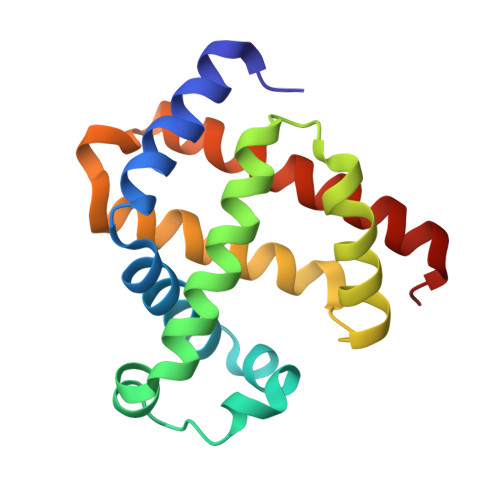
Distal pocket control of nitrite binding in myoglobin
Yi, J., Thomas, L.M., Richter-Addo, G.B.(2012) Angew Chem Int Ed Engl 51: 3625-3627
- PubMed: 22383424
- DOI: https://doi.org/10.1002/anie.201200010
- Primary Citation of Related Structures:
3V2V, 3V2Z
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Oklahoma, 101 Stephenson Parkway, Norman, OK 73019, USA. yijun@ou.edu