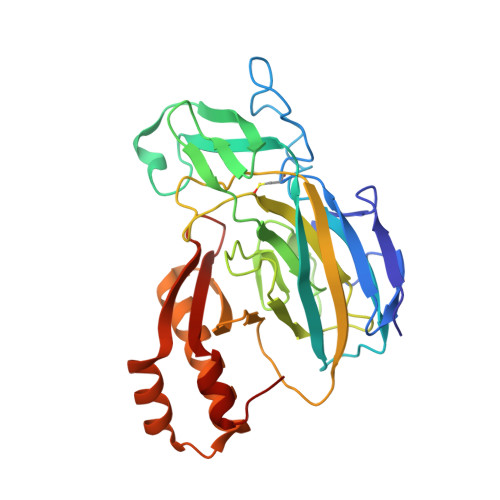
Crystal Structure Analysis of a Recombinant Predicted Acetamidase/Formamidase from the Thermophile Thermoanaerobacter tengcongensis.
Qian, M., Huang, Q., Wu, G., Lai, L., Tang, Y., Pei, J., Kusunoki, M.(2012) Protein J 31: 166-174
- PubMed: 22207484
- DOI: https://doi.org/10.1007/s10930-011-9387-0
- Primary Citation of Related Structures:
3TKK - PubMed Abstract:
The structure of acetamidase/formamidase (Amds/Fmds) from the archaeon Thermoanaerobacter tengcongensis has been determined by X-ray diffraction analysis using MAD data in a crystal of space group P2₁, with unit-cell parameters a = 41.23 (3), b = 152.88 (6), c = 100.26 (7) Å, β = 99.49 (3) ° and been refined to a crystallographic R-factor of 17.4% and R-free of 23.7%. It contains two dimers in one asymmetric unit, in which native Amds/Fmds (TE19) contains of the 32 kDa native protein. The final model consists of 4 monomer (299 amino acids residues with additional 2 expression tag amino acids residues), 5 Ca²⁺, 4 Zn²⁺ and 853 water molecules. The monomer is composed by the following: an N-domain which is featuring by three-layers β/β/β; a prominent excursion between N-terminal end of strand β₇ and β₁₁, which contains four-stranded antiparallel β sheet; an C-domain which is formed by the last 82 amino acid residues with the feature of mixed α/β structure. The protein contains ion-pair Ca²⁺-Zn²⁺. The portion of three-layer β/β/β along with the loops provides four protein ligands to the tightly bound Ca²⁺, three water molecules complete the coordination; and provides five protein ligands to the tightly bound Zn²⁺, one water molecule complete the coordination.
Organizational Affiliation:
State Key Laboratory for Structural Chemistry of Unstable and Stable Species, College of Chemistry, Peking University, Beijing, People's Republic of China. qianmx@pku.edu.cn