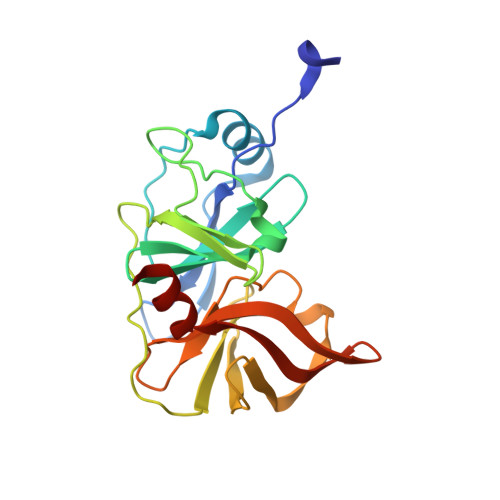
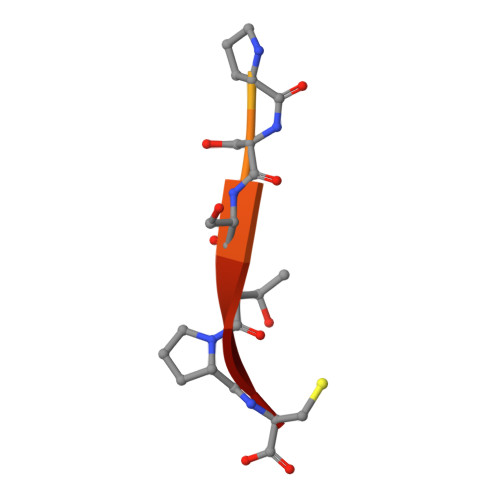
Molecular mechanisms of viral and host cell substrate recognition by hepatitis C virus NS3/4A protease.
Romano, K.P., Laine, J.M., Deveau, L.M., Cao, H., Massi, F., Schiffer, C.A.(2011) J Virol 85: 6106-6116
- PubMed: 21507982
- DOI: https://doi.org/10.1128/JVI.00377-11
- Primary Citation of Related Structures:
3RC4, 3RC5, 3RC6 - PubMed Abstract:
Hepatitis C NS3/4A protease is a prime therapeutic target that is responsible for cleaving the viral polyprotein at junctions 3-4A, 4A4B, 4B5A, and 5A5B and two host cell adaptor proteins of the innate immune response, TRIF and MAVS. In this study, NS3/4A crystal structures of both host cell cleavage sites were determined and compared to the crystal structures of viral substrates. Two distinct protease conformations were observed and correlated with substrate specificity: (i) 3-4A, 4A4B, 5A5B, and MAVS, which are processed more efficiently by the protease, form extensive electrostatic networks when in complex with the protease, and (ii) TRIF and 4B5A, which contain polyproline motifs in their full-length sequences, do not form electrostatic networks in their crystal complexes. These findings provide mechanistic insights into NS3/4A substrate recognition, which may assist in a more rational approach to inhibitor design in the face of the rapid acquisition of resistance.
Organizational Affiliation:
University of Massachusetts Medical School, Department of Biochemistry and Molecular Pharmacology, 364 Plantation Street, Worcester, MA 01605, USA.